Python integration
GA3 Python nodes support a wide range of Python packages, enabling advanced data analysis, visualization, and computational capabilities. By integrating scientific libraries directly into GA3 recipes, the platform’s functionality can be extended to meet specific research needs. Whether performing statistical analysis, processing images, analyzing large datasets, or applying machine learning techniques, Python nodes offer the flexibility to enhance workflows with customized scripts.
This section provides examples of how various scientific Python packages can be used within GA3, helping to streamline research and maximize the available tools.
For more GA3 examples, check the Laboratory Imaging GitHub repository GA3-examples.
NIS-Express comes with many scientific packages preinstalled. In order to install additional python modules use separate environments or managed environment in the Python node (see below).
NumPy for simple threshold
This workflows showcases a simple working example of using NumPy library for segmentation (see numpy.org).
Setup
Open the file that needs to be segmented.
In NIS Express GA3 Editor:
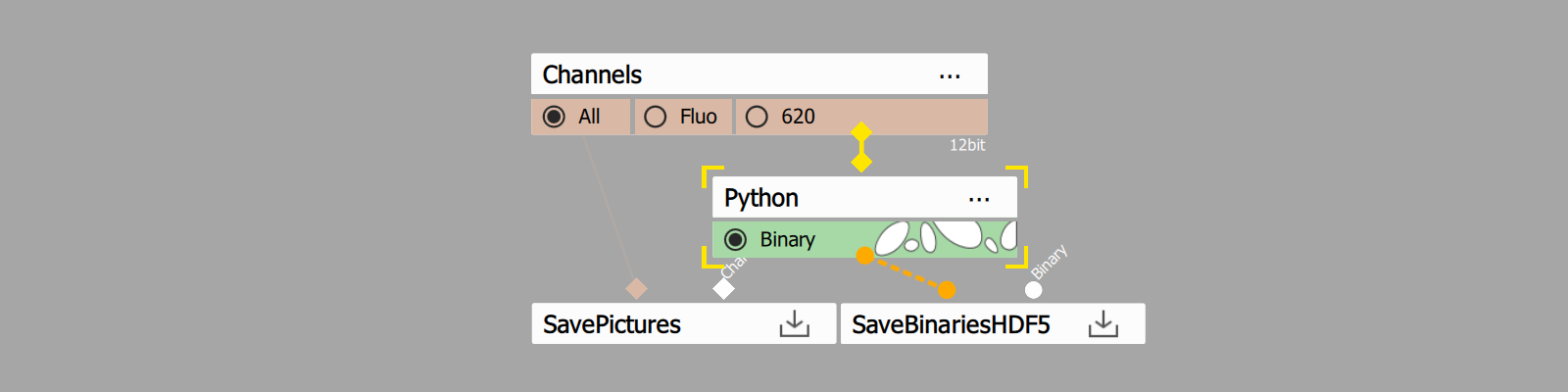
- Add Python node and open its settings
- Add color input
- Add binary output
- Insert code from Code section.
- Connect the input pin to the Channels output to be segmented.
- Connect the output pin to the SaveBinariesHDF5 node.
Code
| |
The code:
- Imports NumPy. [
line 3] - Names binary output as “Binary” and sets its color green. [
line 7] - Get color data. [
line 15] - Segments values in range from 300 to 400 (edit as needed) [
line 16] - Stores data to output as unsigned 8bit integer. [
line 17]
Results
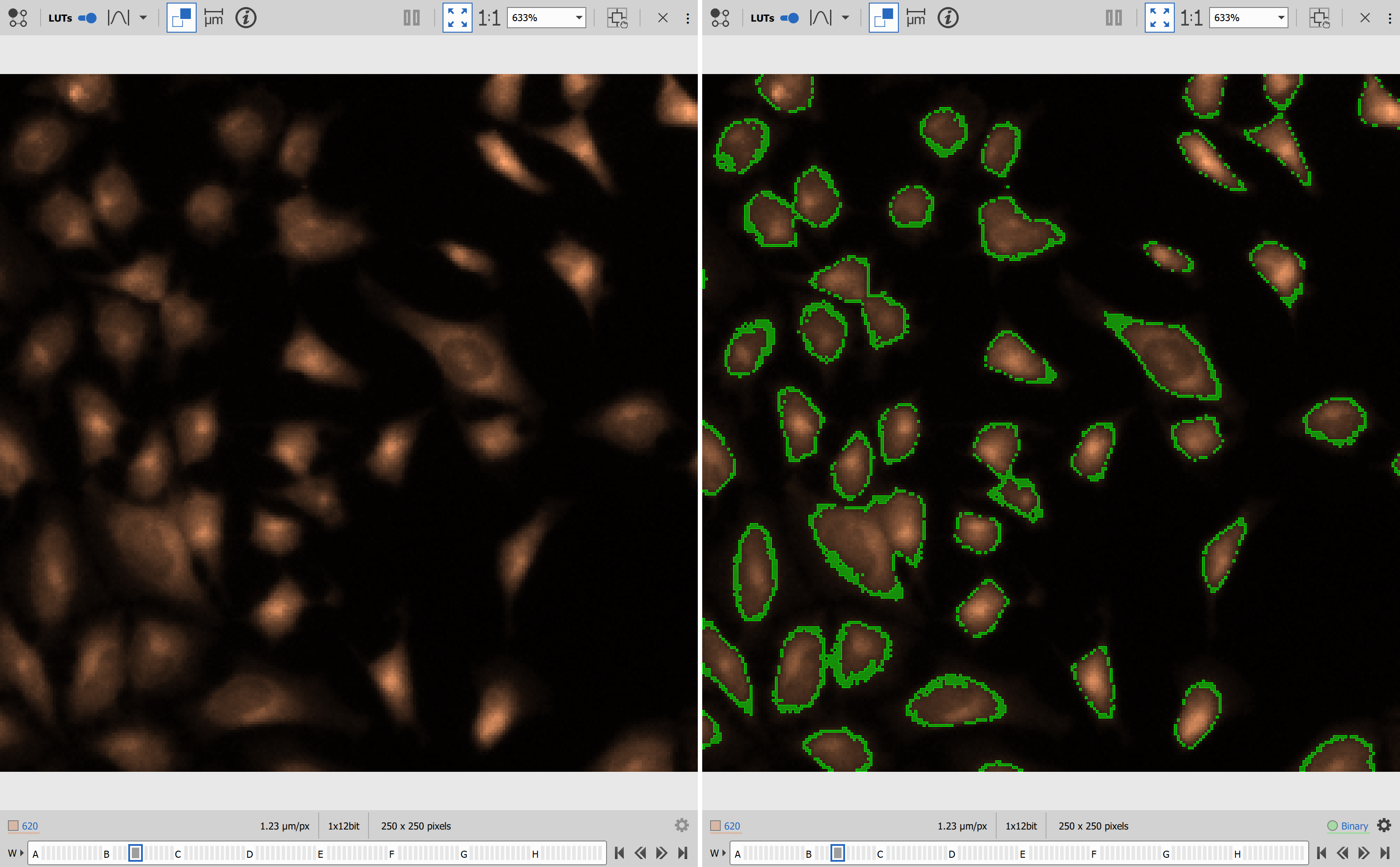
scikit-image for segmentation
Scikit-image is “a collection of algorithms for image processing” (see scikit-image.org).
Setup
Open the file that needs to be segmented.
In NIS Express GA3 Editor:
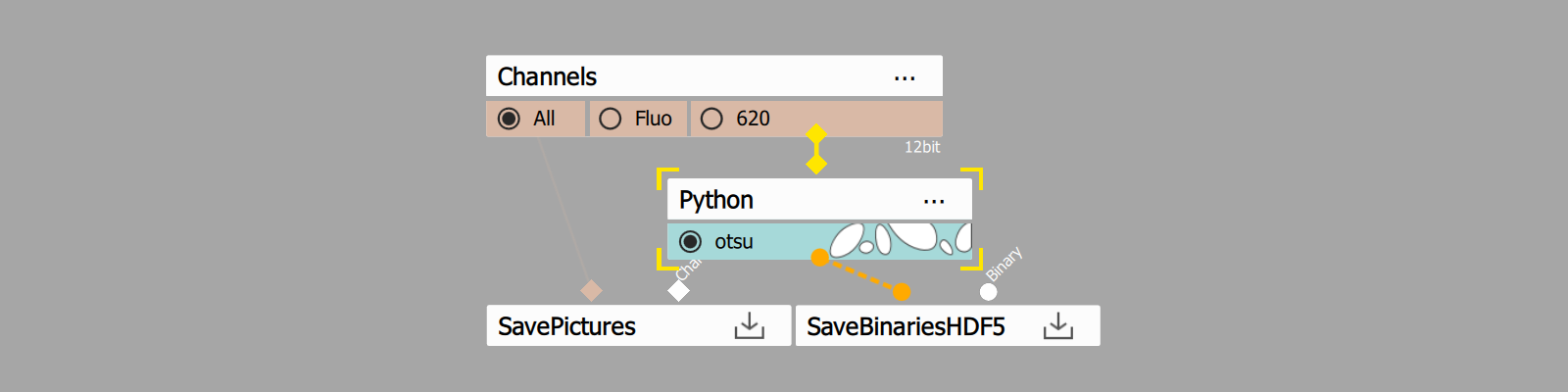
- Add Python node and open its settings
- Add color input
- Add binary output
- Insert code from Code section.
- Connect the input pin to the Channels output to be segmented.
- Connect the output pin to the SaveBinariesHDF5 node.
Code
| |
- Imports NumPy and filters from the scikit-image package [
lines 2, 3]. - Defines the output to be a new binary named “otsu” and sets cyan color. [
line 7]. - Takes the image from the inp array [
line 15]. - Calculates the threshold calling threshold_otsu [
line 16]. - Creates the binary by directly comparing the values from the image with the threshold value [
line 17]. - Sets the binary data into the out array while converting to the proper format (uint8 or int32 for binary IDs) [
line 18].
Results
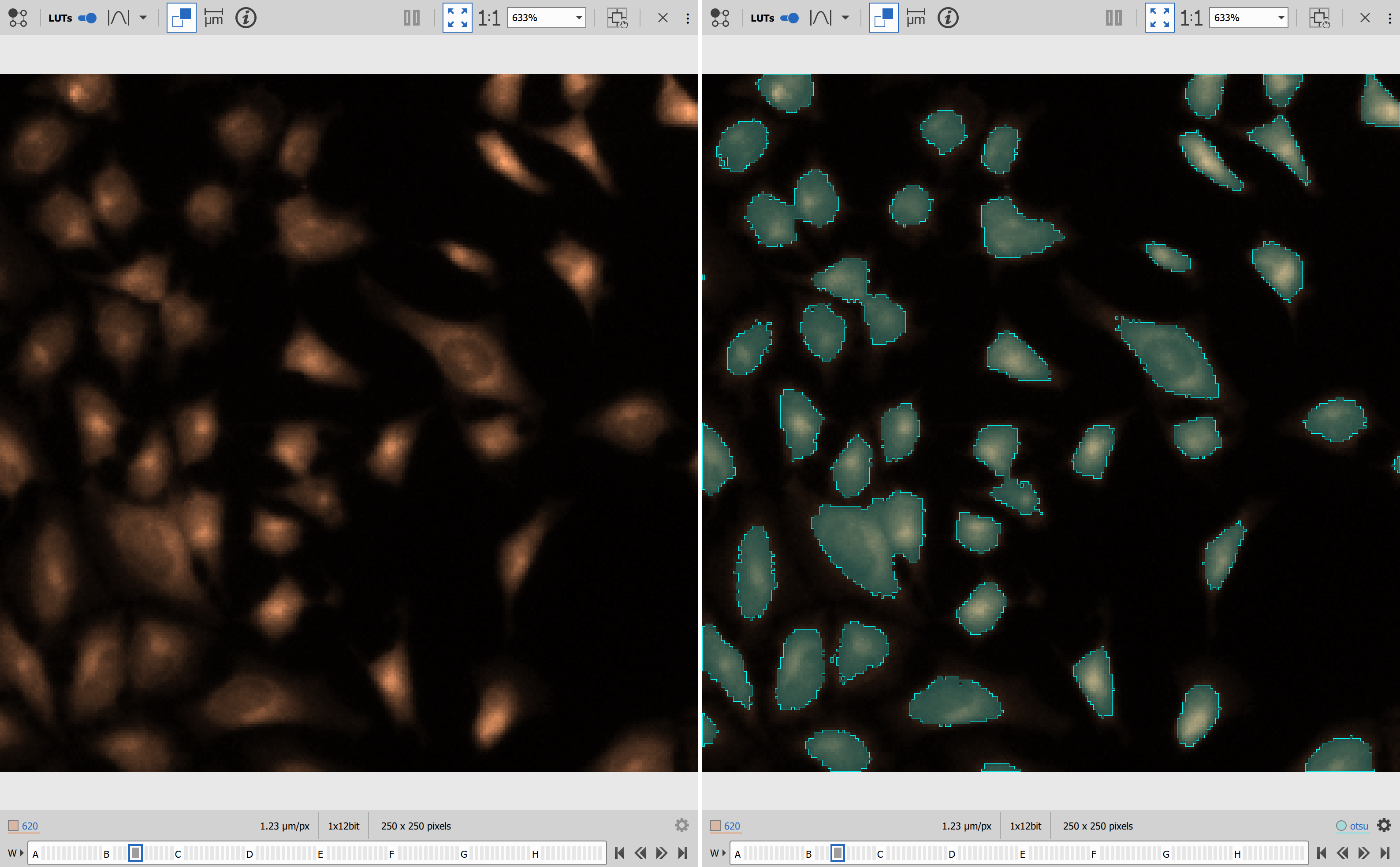
BaSiCPy for shading removal
BaSiCPy is a library “for background and shading correction of optical microscopy images” (see BaSiCPy).
Setup
Open the file that needs to be processed.
In NIS Express GA3 Editor:
- Add Channel input.
- Add Channel output.
- Set the python mode to “Managed environment”
- Insert the environment from the Environment section.
- Insert the code from Code section.
- Connect the input pin to the Channels output to be processed.
- Connect the output pin to the SavePictures node.
Environment
channels:
- conda-forge
dependencies:
- python=3.10
- pip
- pip:
- "scipy<1.13"
- "jax[cpu]==0.4.13"
- "hyperactive<5"
- "ml-dtypes==0.2.0"
- basicpy
variables:
PIP_FIND_LINKS: "https://whls.blob.core.windows.net/unstable/index.html"
PIP_USE_DEPRECATED: "legacy-resolver"The environment was created and tested in the November, 2025.
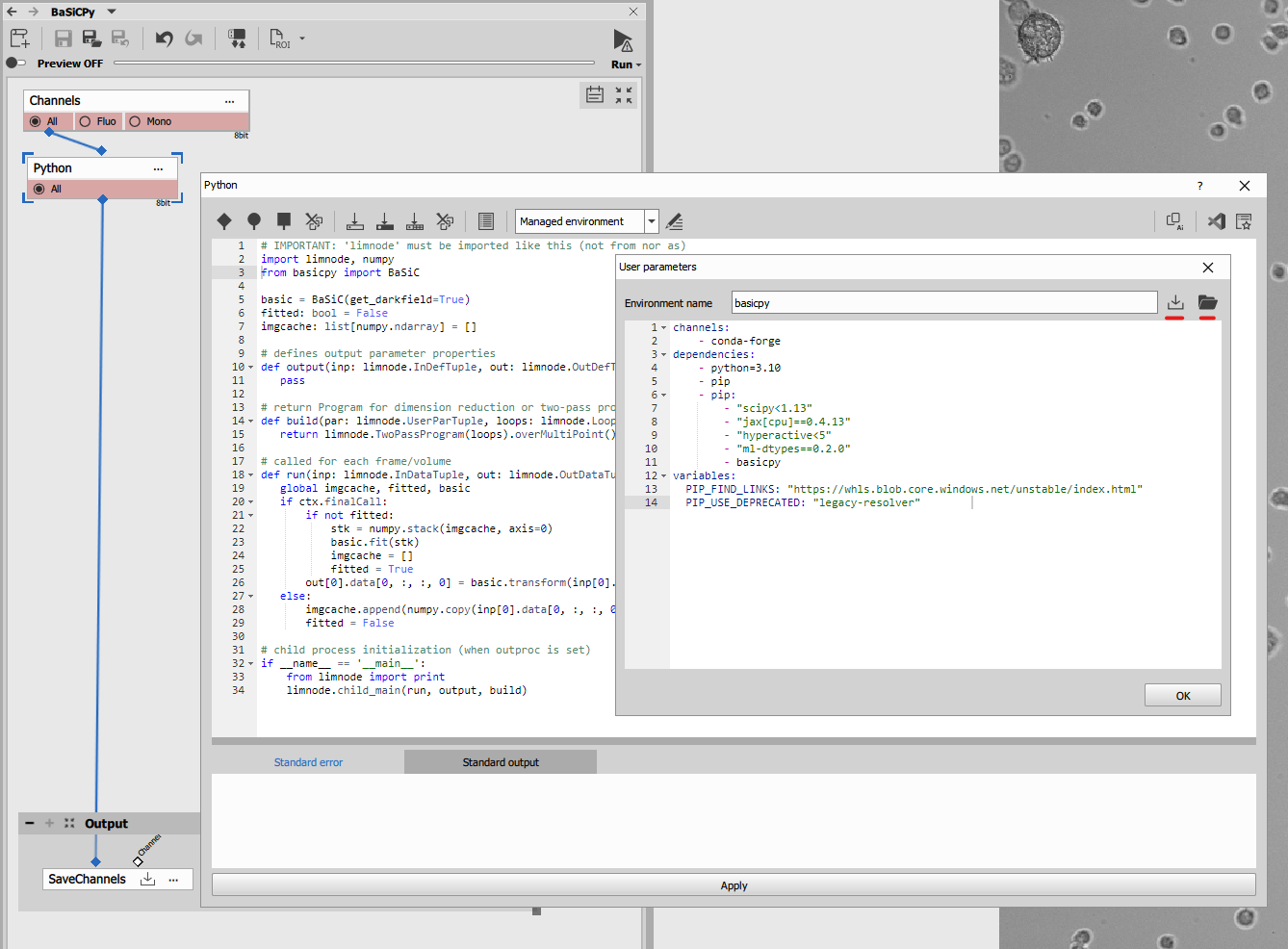
Code
| |
- Imports NumPy and basicpy from the BaSiC package [
lines 2, 3]. - Initializes global variables for two pass processing [
lines 5-7]. - Sets up two pass progam [
line 15]. - Enables using global variables in run [
line 19]. - Calculates shading correction (only once) and applies to every frame in second pass[
lines 21-26]. - Collects data in first pass [
lines 27-29].
Results
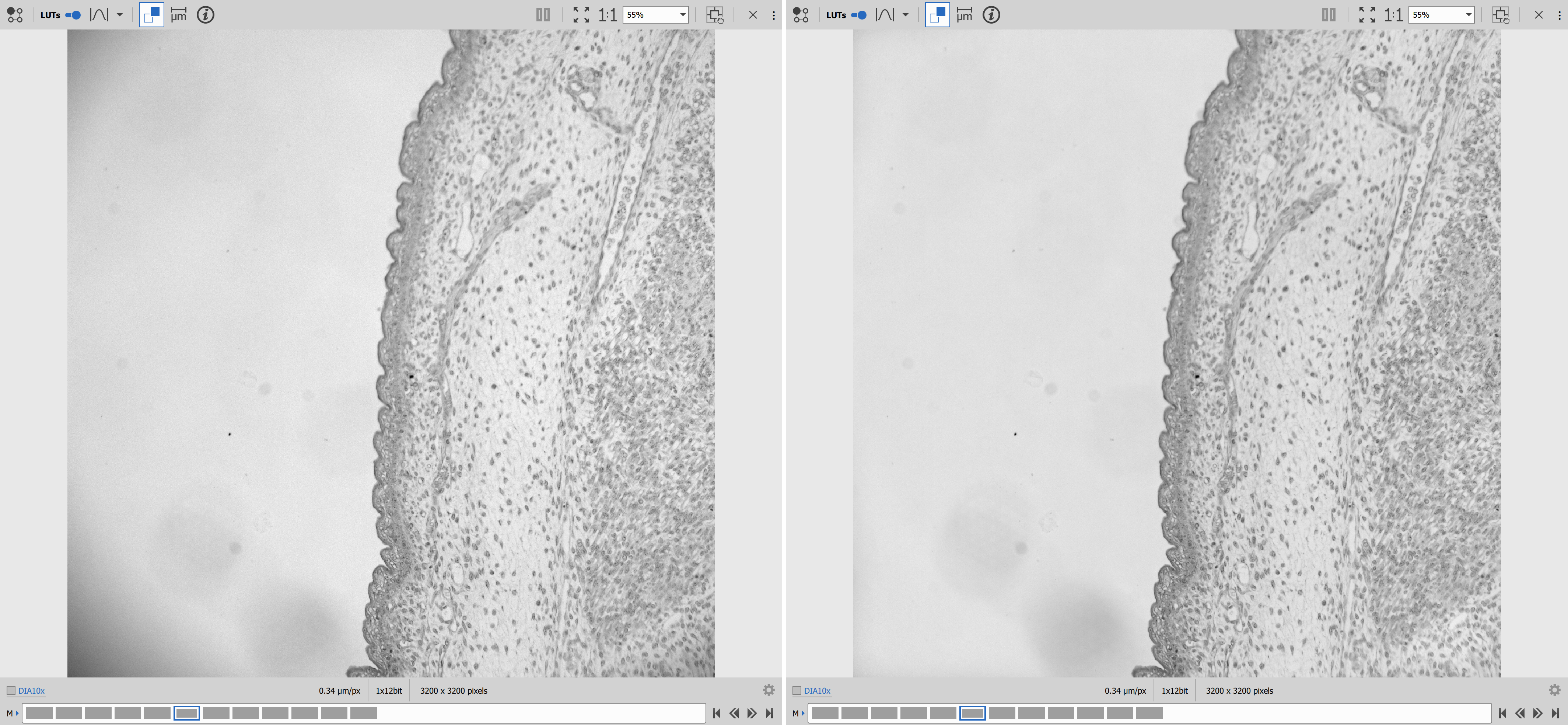
Matplotlib for data visualization
Matplotlib comes with NIS-Express.
This example will built on and modify the Object counting example.
Open the
02_count_in_t.nd2time-lase nd2 image and02_object_count_fluo.ga3recipe.
Replace both graph nodes with a Python node. Both having Table as single input and single output.
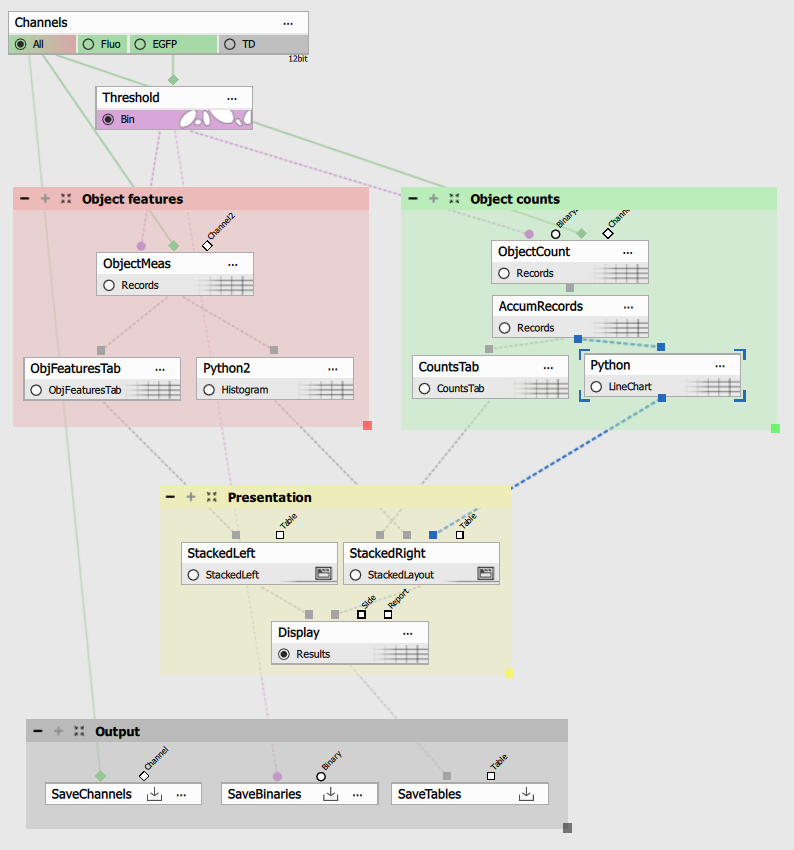
Object count vs. time
- In the
output()function [line 37] make the table result and provide it with input parameter (for taking the accumulation from theinp[0]), - define the
draw_graph()function [line 19] that actually draws the graph giving it the x, y data and background and foreground colors, - call the draw_graph function twice for generating light and dark graphs [
lines 48, 49] and finally - call the
withMplImage()method [line 52] to render the matplotlib figure into an image.
The withMplImage() takes following arguments:
- a tuple with ND loop indexes of the current frame or accumulated loop
(
ctx.inpParameterCoordinates[i]whereiis the index of the input parameter is a good starting point) and - a single figure or pyplot object or tuple of two such objects for light and dark color scheme.
matplotlib.pyplot global instance which may be in
unexpected state when the node run function is called. | |
The above python code produces following line chart.
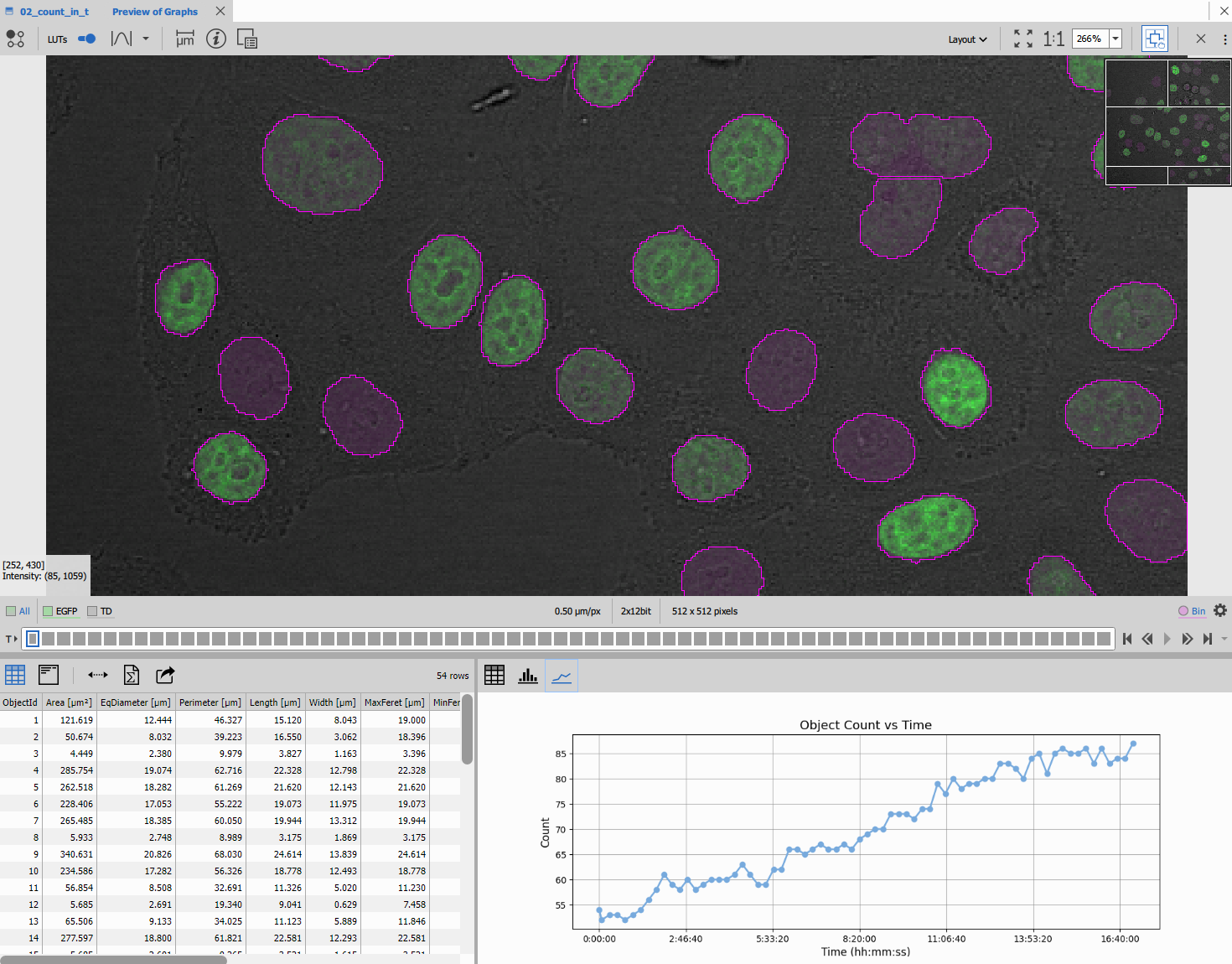
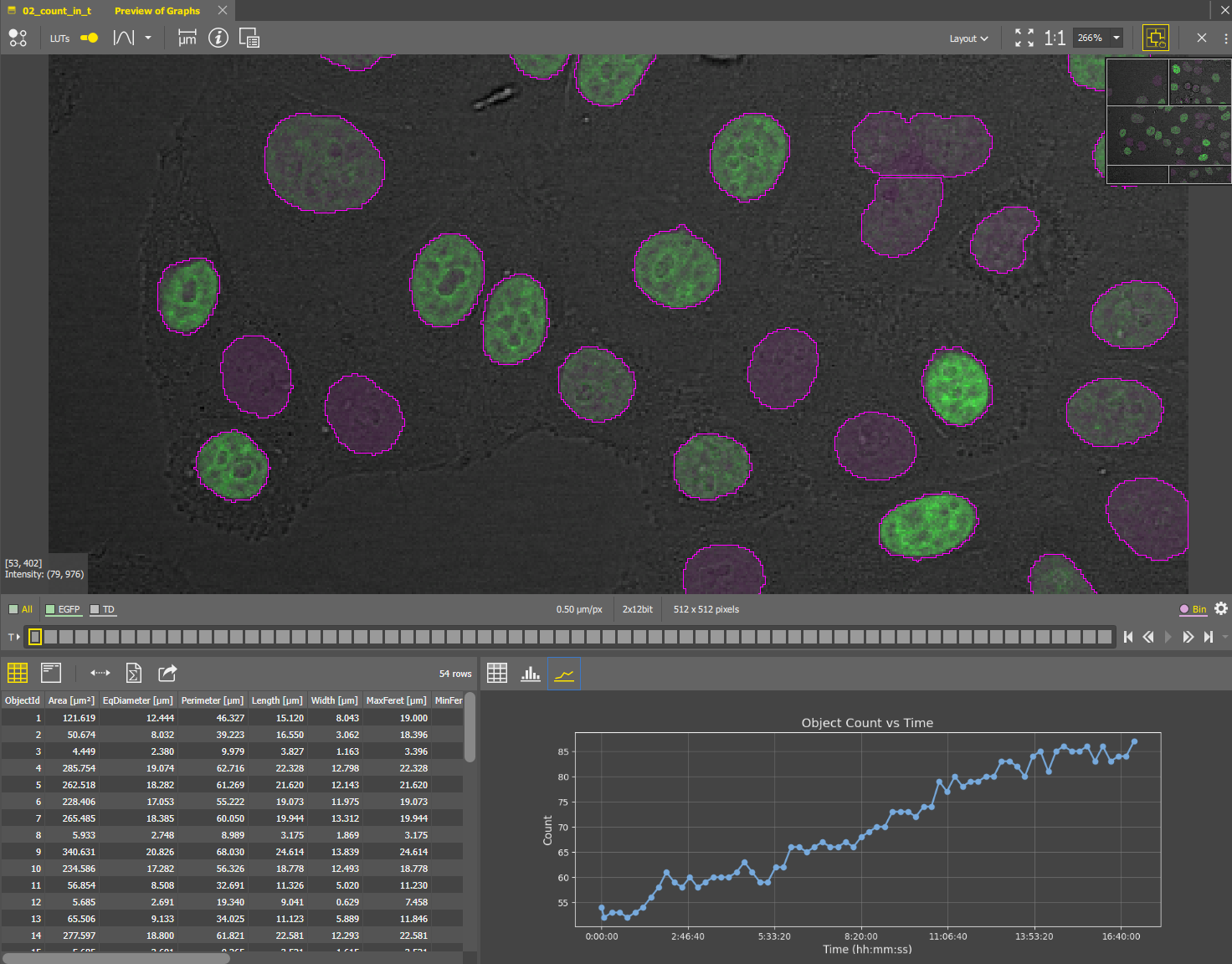
Histogram of object sizes per frame
Similarly with histogram:
| |
The above python code produces following histogram.
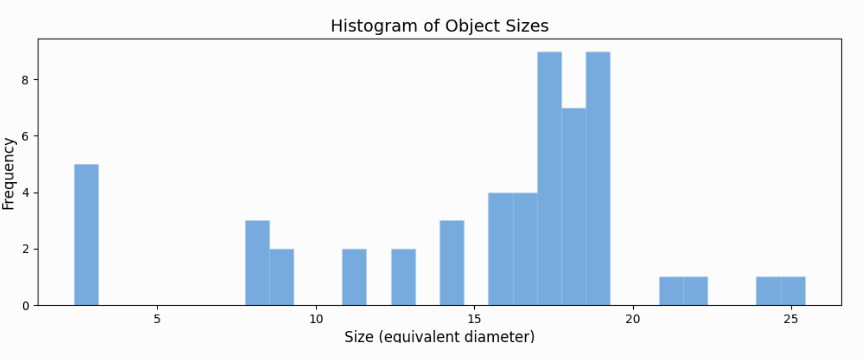
Conclusion
These two examples are not the fanciest plots. The lack interactivity as the native GA3 plots because they are rendered into a static image (or two: light and dark). However based on the aggregation of the inputs and loop coordinates the histogram will change as the current frame changes and linechart of counts vs. time will not as there is only one plot of aggregated counts.
Omnipose for bacteria segmentation
Omnipose is a pupular general image segmentation package that builds on Cellpose.
Setup
In the GA3 node add
- one channel input,
- one binary output,
- one channel output and
- set the Execution mode to “Managed environment”
- insert the environment from the Environment section.
- insert the code from Code section.
- connect the input pin to the Channels output to be processed.
- connect the binary output pin to the SavePictures node.
- connect the channel output pin to the SaveBinaries node.
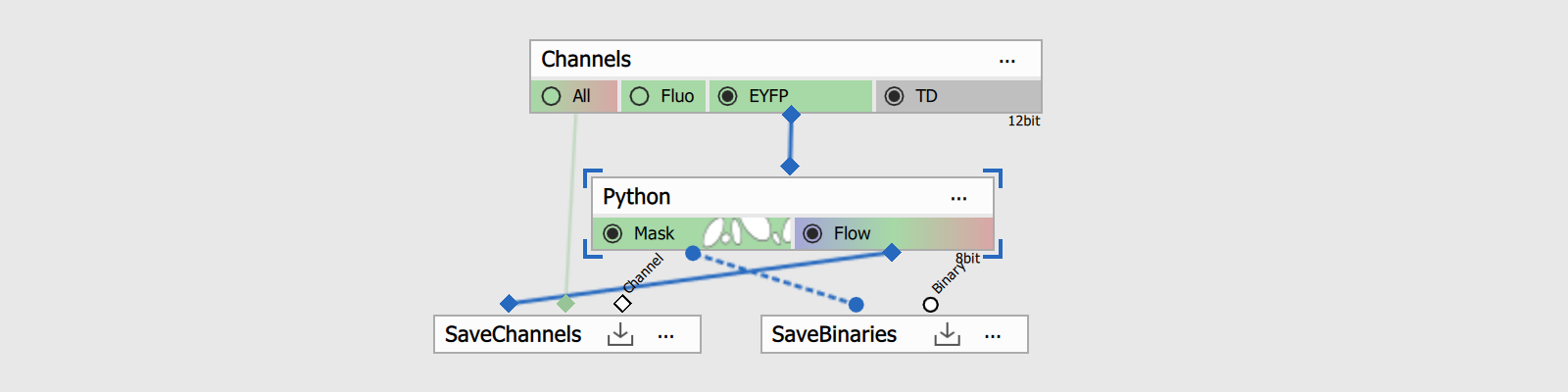
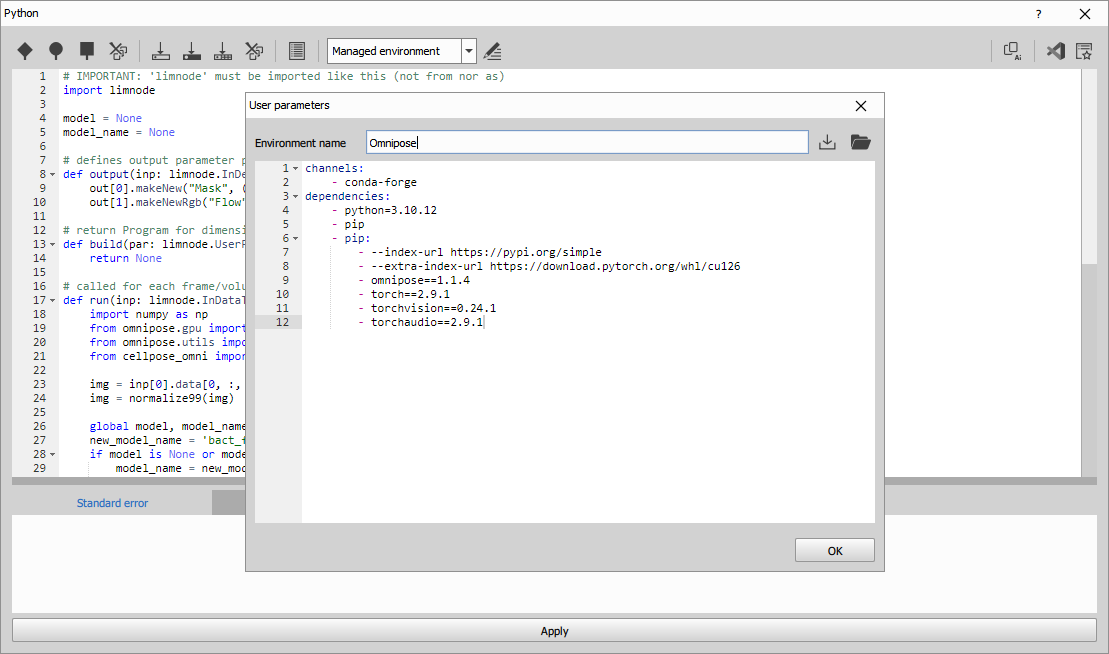
Environment
Edit the evironment and insert following definition:
channels:
- conda-forge
dependencies:
- python=3.10.12
- pip
- pip:
- omnipose==1.1.4channels:
- conda-forge
dependencies:
- python=3.10.12
- pip
- pip:
- --index-url https://pypi.org/simple
- --extra-index-url https://download.pytorch.org/whl/cu126
- omnipose==1.1.4
- torch==2.9.1
- torchvision==0.24.1
- torchaudio==2.9.1Then install it by clicking the “Install managed environment” icon. Be patient it takes time.
The environment was created and tested in the January, 2026.
Code
Insert the code as follows:
| |
The code:
- Initializes the global variable
modelso the model is created only once. [line 4] - Initializes the global variable
model_nameto track the currently selected model type. [line 5] - Defines a binary output named “Mask” and sets its display color to green. [
line 8] - Defines a channel output named “Flow” and sets its format to 8-bit RGB. [
line 9] - Checks whether a GPU device is enabled [
line 30] - Creates the CPU model. [
line 32] - Creates the GPU model. [
line 34] - Runs the model on the current frame using the specified parameters. [
line 52] - Splits the labeled mask into separate binary objects. [
line 53] - Writes the resulting binary mask to the binary output [
line 54] - Applies the binary mask to the flow image, converts RGB to BGR, and writes the result to the channel output. [
line 55]
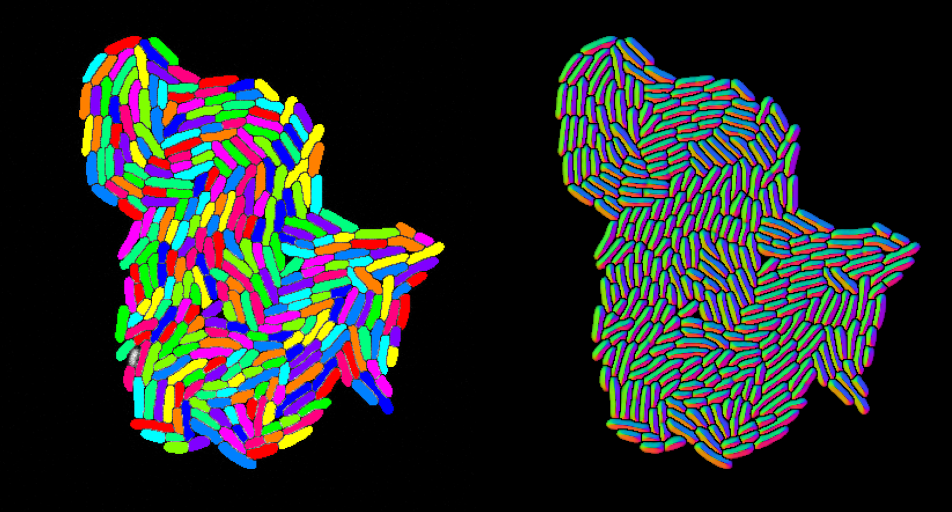
Results

Export table to CSV
This workflow demonstrates a simple table export to CSV with customizable export options using an underlying pandas DataFrame (see pandas.pydata.org). It also shows how to add parameters for User Mode configuration.
Setup
Open the file that needs to be segmented.
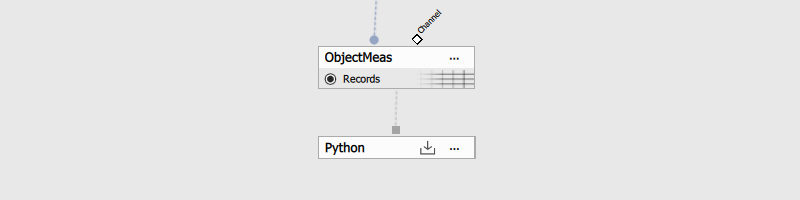
In NIS Express GA3 Editor:
- Add Python node and open its settings
- Add table input
- Insert code from Code section.
- Connect the input pin to the Table output to be exported.
Code
| |
The code:
- Imports
Pathfrompathlib[line 3] - Reads GUI parameters [
lines 7-9] - Extracts output image filepath info for use as default [
lines 19-21] - Resolves output path, adds default CSV name if missing, and normalizes it [
lines 23-28] - Extracts final filename, extension, and target directory [
lines 30-32] - Creates the output directory if it does not exist [
line 34] - Appends loop indices to filename to ensure unique files per iteration. [
lines 36-38] - Exports the input table to CSV with selected separator and decimal format [
line 40]
User parameters
- Open the
User parametersin the top toolbar of the node. - In the opened window:
- fill the description:
Select options for CSV export - add parameters
- Output
- Type:
str - Definition:
{"control":"Path","mode":"save","type":"file"}
- Type:
- Delimiter
- Type:
str - Initial value:
; - Definition:
{"control":"Selection","list":[{"text":"Semicolon ';'","value":";"},{"text":"Comma ','","value":","},{"text":"Tab '\\t'","value":"\t"}]}
- Type:
- Decimal separator
- Type:
str - Initial value:
. - Definition:
{"control":"Selection","list":[{"text":"Dot '.'","value":"."},{"text":"Comma ','","value":","}]}
- Type:
- Output
- fill the description:
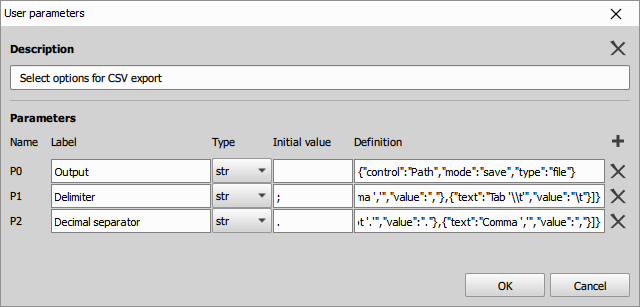
- Close the
User parameterswindow - Switch to the
User modein the top toolbar of the node. - In the newly opened window click “Skip”
- Window should switch to User mode.
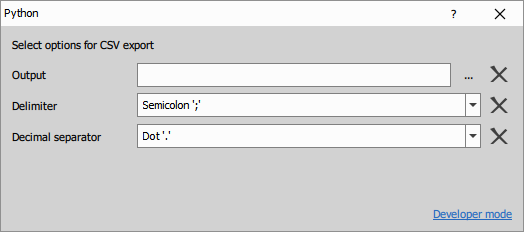
- If needed, return back by clicking Developer mode.
Results
CSV files are exported to the selected output directory, and export options can be easily modified via the custom GUI.