Object counting
Open the workflow in NIS-Express:
Object counting is the most fundamental measurement workflow in GA3. At its core, it requires only two steps: segmentation to detect objects and object count measurement to quantify them.
However, this workflow demonstrates a complete analysis pipeline that produces comprehensive results including temporal trends, per-object features, and synchronized visualizations.
What this workflow produces
This workflow creates a complete analysis package:
- Per-field statistics table: Object count, total area, and area fraction for each frame in the time-lapse
- Per-object features table: Morphological measurements for every detected object in the current frame
- Temporal trend graph: Line chart showing how object count and area fraction evolve over time
- Feature distribution histogram: Shows the distribution of any selected object feature
- Synchronized interface: Links all visualizations to the image, enabling interactive exploration
The result provides both population-level statistics (field counts over time) and individual-level data (per-object measurements).
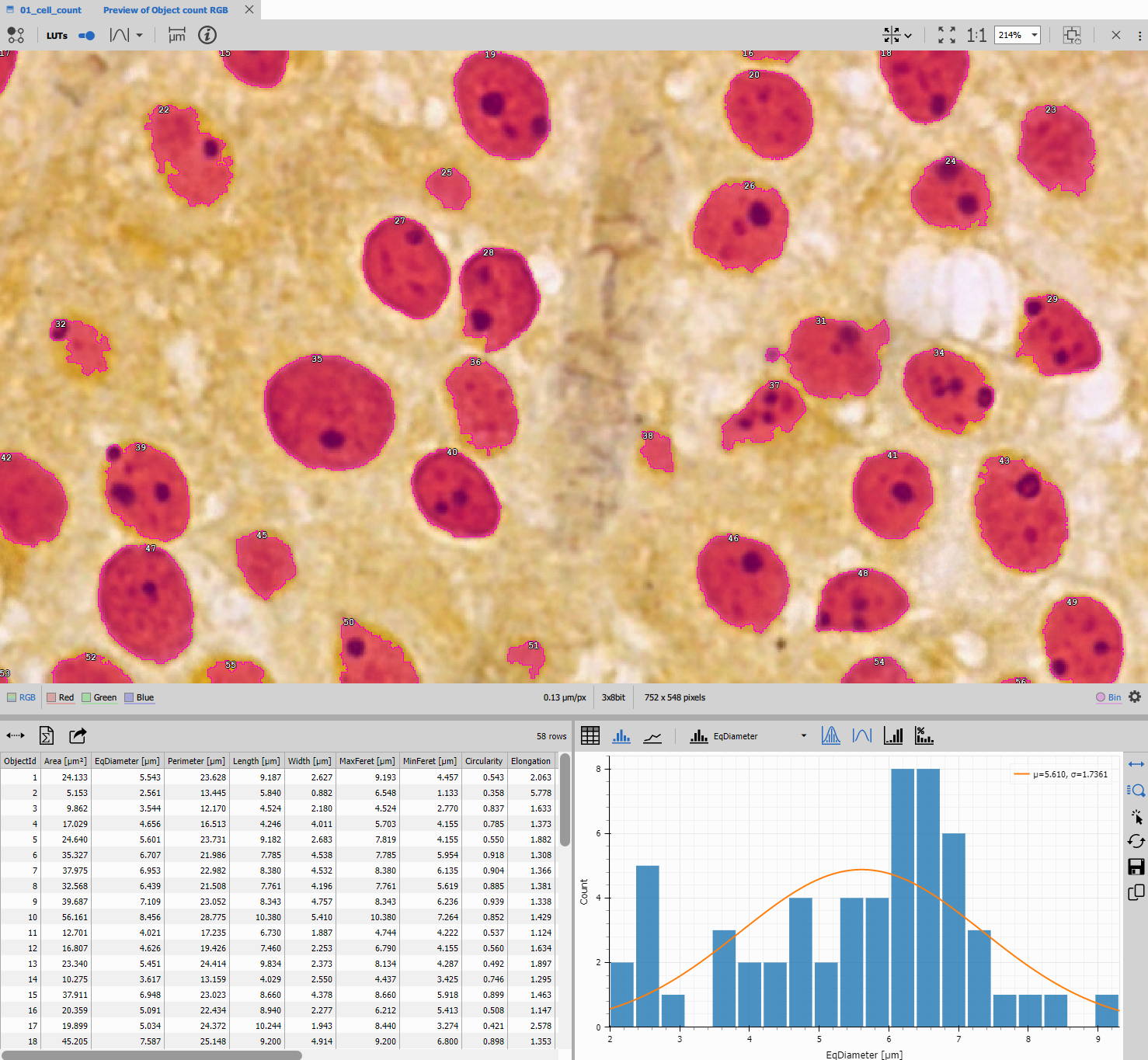
Workflow overview
The recipe consists of four functional blocks:
Segmentation
Detects objects using thresholding (or alternative segmentation methods).
Object counting block
Measures field-level statistics, accumulates data across time, and displays temporal trends.
Object measurement block
Measures detailed features for individual objects in the current frame.
Presentation block
Organizes all tables and graphs into a synchronized, interactive display.
Recipe variations
This workflow comes in three variations that share the same core structure:
RGB version (single frame):
- Uses RGB Threshold for color images
- Processes a single RGB image
- Demonstrates the basic workflow structure
Fluorescence version (time-lapse):
- Uses Threshold for fluorescence channels
- Processes multi-frame time-lapse data
- Shows temporal evolution of object counts
Catalog version (with gallery):
- Adds Object Catalog node
- Displays a grid gallery of detected objects
- Includes border object removal for cleaner results
---
config:
look: handDrawn
theme: neutral
---
graph TD
subgraph "Object counting"
objcount[Object Count]
accum[Accumulate Records]
countstab[Table - All Frames]
linechart[Line Chart]
end
subgraph "Object measurement"
objmeas[Object Features]
objtab[Table - Current Frame]
histo[Histogram]
end
subgraph "Presentation"
left[Stacked Layout - Left]
right[Stacked Layout - Right]
display[Display]
end
subgraph Outputs
savechn[Save Channels]
savebin[Save Binaries]
savetab[Save Tables]
end
%% Main workflow
channels[Channels] -->|RGB or Fluo| threshold[Threshold] -->|Bin| objcount
channels -.->|All| objcount
threshold -->|Bin| objmeas
channels -.->|All| objmeas
%% Object counting flow
objcount -->|Records| accum
accum -->|Records| countstab
accum -->|Records| linechart
%% Object measurement flow
objmeas -->|Records| objtab
objmeas -->|Records| histo
%% Presentation
objtab --> left
countstab --> right
histo --> right
linechart --> right
left --> display
right --> display
%% Outputs
channels -->|All| savechn
threshold -->|Bin| savebin
display -->|Results| savetab
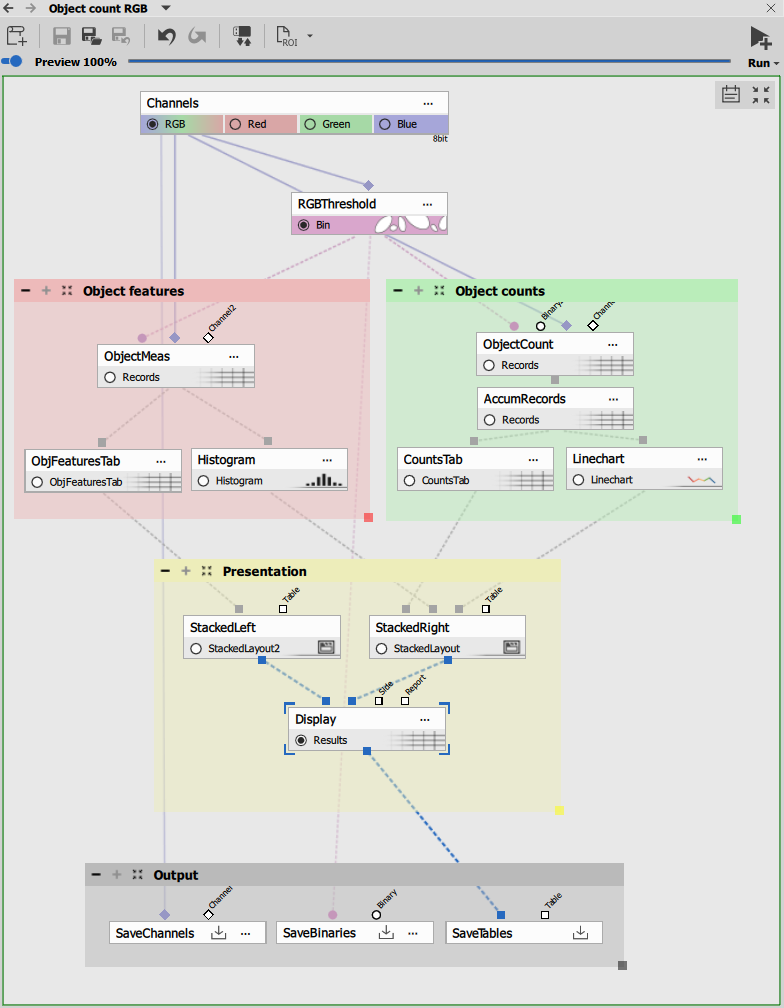
Segmentation
Segmentation is the critical first step that determines which pixels belong to objects of interest. GA3 provides multiple segmentation methods, with thresholding being the most straightforward.
Threshold methods
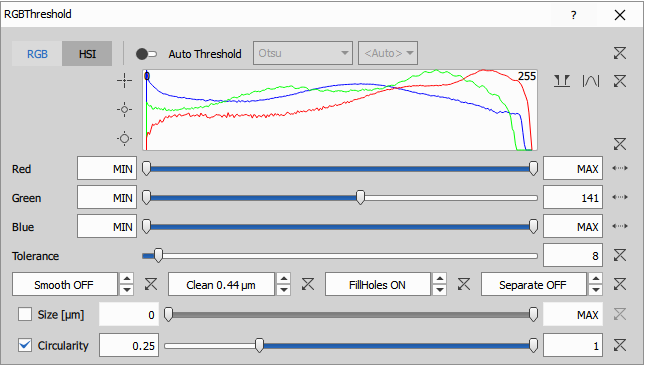
For RGB images - RGB Threshold:
- Works directly on RGB color images
- Separates objects based on color (hue, saturation, intensity)
- Settings used:
- Auto Threshold Method: Intermodes algorithm
- IsHSI: 1 (uses HSI color space instead of RGB)
- GreenHigh: 142 (upper limit on green channel in HSI)
- Clean: 0.504 µm to remove small debris
- Smooth: 0.063 µm to regularize boundaries
- Fill: Enabled to fill holes inside objects
- FilterByCircularity: Enabled with CircularityLow: 0.25 to remove elongated artifacts
For fluorescence images - Threshold:
- Works on grayscale fluorescence channels
- Separates bright objects from dark background
- Settings used:
- Auto Threshold Method: Triangle algorithm
- AutoThresholdOn: Enabled for automatic threshold determination
- Clean: 0.498 µm to remove noise
- Separate: 0.498 µm to split touching objects
- Smooth: 0.498 µm to smooth boundaries
- Fill: Enabled to fill holes
- IntensityLow: 114 (manual override if needed)
Auto threshold algorithms: GA3 provides many automatic thresholding methods (Triangle, Intermodes, Otsu, Li, etc.). Each works best for different intensity distributions. If auto settings don’t work well:
- Try different auto methods from the dropdown
- Use the histogram to manually set IntensityLow/High
- Check the preview to verify segmentation quality
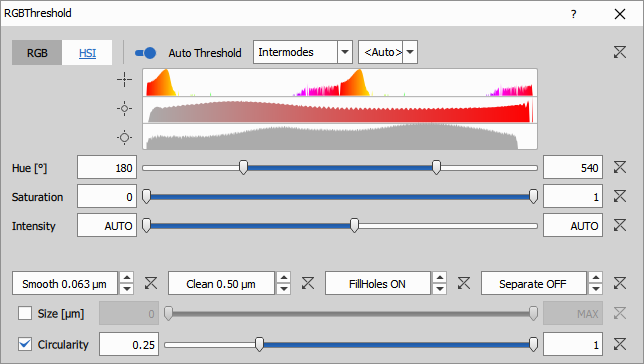
Alternative segmentation methods
While thresholding works for many samples, GA3 offers more sophisticated segmentation for challenging cases:
- For small, bright, round objects (cells, particles, nuclei)
- Uses contrast and expected diameter
- More robust to background variations
- AI-powered segmentation for complex structures
- Learns from training data
- Handles variable morphology
- AI segmentation for specific object types
- Pre-trained models available
- Best for cells, nuclei, organoids
- State-of-the-art cell segmentation
- Works without training data
- Excellent for cells and nuclei
Choose the segmentation method that best matches your sample characteristics.
Object counting block
This block measures field-level (population) statistics and tracks them across time. It produces a table showing how object counts change over the time-lapse sequence.
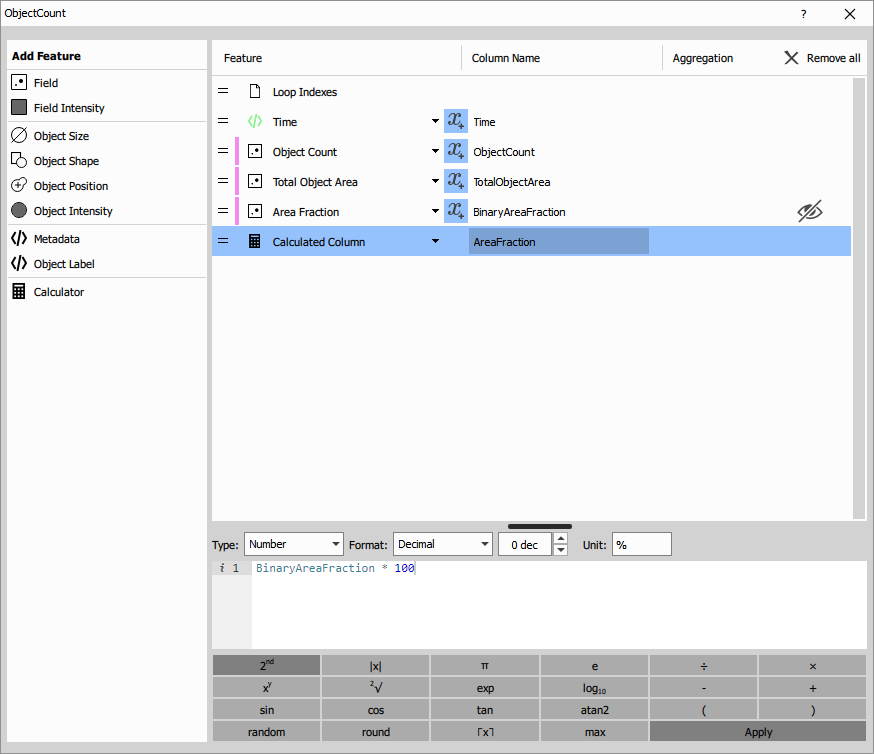
Object Count measurement
Object Count computes aggregate statistics for all objects in each field. Unlike measuring individual objects, this node produces one row per frame containing population-level data.
Measured features:
- Time: Timestamp of the frame (seconds or frame number)
- Object Count: Total number of detected objects in the field
- Total Object Area: Sum of all object areas (µm²)
- Area Fraction: Proportion of the field covered by objects (0-1)
Calculated feature:
- Area Fraction %: Calculated column =
AreaFraction × 100for easier interpretation as percentage
The node has two inputs:
- Binary layer: The segmentation result defining objects
- Channel: For intensity measurements in the objects
This measurement runs frame-by-frame through the time-lapse, producing one row per frame.
Accumulate Records
Accumulate Records collects measurements from all frames into a single table. Without accumulation, only the current frame’s data would be available.
Key settings:
- LoopName: “All loops” (accumulate over all loops - time-lapse, z-stack, multi-point)
- PreviewMode: OFF (accumulate all data in preview)
The accumulated table contains one row per frame, allowing temporal analysis of the entire sequence.
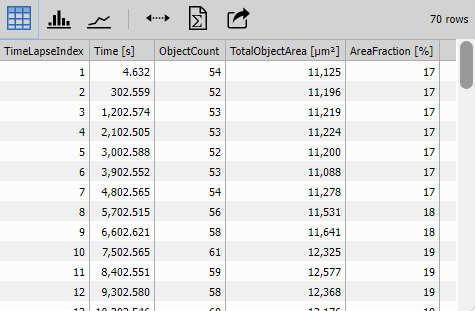
Counts Table
Table displays the accumulated object counts with interactive functionality.
Configuration:
- Data/Display: “All data” to show all frames
- Interaction/Row selection: Enabled to link table rows to image frames
- Statistics: Shows min, max, mean of all columns
Interactive behavior:
- Clicking a row in the table jumps to that frame in the image
- The current frame is highlighted in the table
- Enables quick navigation through the time-lapse
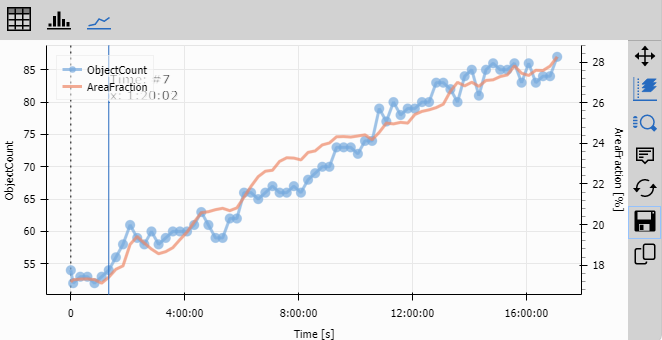
Line Chart
Line Chart visualizes the temporal evolution of object populations.
Configuration:
- Data/Display: “All data” to plot all frames
- X-axis: Time (with format set to display as time)
- Left Y-axis: Object Count
- Right Y-axis: Area Fraction % (secondary axis for different scale)
Display features:
- Dual Y-axes allow plotting variables with different scales
- Time formatting displays timestamps properly
- Interactive tools: pan, zoom, frame selection
This chart immediately reveals trends like:
- Increasing object counts (proliferation, accumulation)
- Decreasing counts (cell death, removal)
- Periodic patterns (division cycles)
- Sudden changes (experimental interventions)
Object measurement block
While the counting block provides population statistics, this block measures detailed features for every individual object. It produces rich morphological and intensity data for single-object analysis.
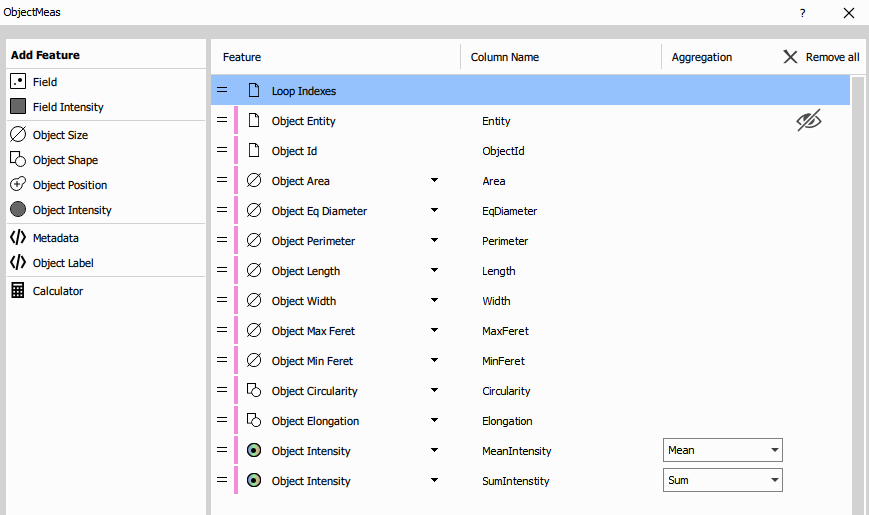
Object Features measurement
Object Features measures comprehensive features for each detected object. This node produces one row per object, with many measured columns.
Inputs:
- Binary layer: Defines object boundaries
- Channel(s): For intensity measurements
Measured feature categories:
Morphological features:
- Size: Area, Equivalent Diameter, Perimeter
- Shape: Length, Width, MaxFeret, MinFeret
- Form factors: Circularity, Elongation
- Position: CenterX, CenterY (object centroids)
Intensity features (per channel):
- MeanIntensity: Average pixel intensity in the object
- SumIntensity: Total intensity (integrated)
- MinIntensity, MaxIntensity: Intensity range
- StdDevIntensity: Intensity variation
This measurement runs per frame, producing a table of objects for the currently displayed frame.
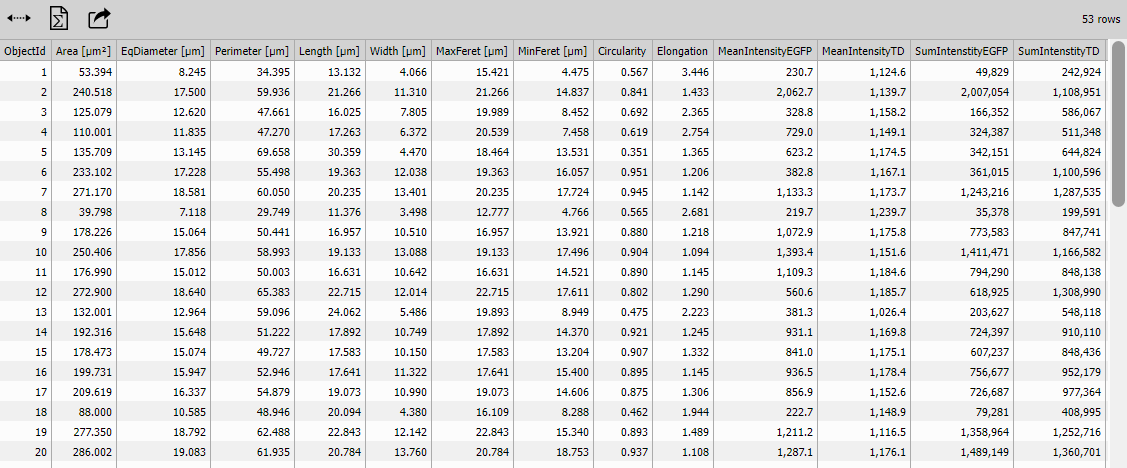
Object Features Table
Table displays individual object measurements.
Configuration:
- Data/Display: “Current frame” to show only objects in the displayed frame
- Interaction/Row selection: Enabled to highlight objects
- Column Sorting: Enabled for sorting by any feature
Interactive behavior:
- Selecting a row highlights the corresponding object in the image
- Selecting an object in the image highlights its row in the table
- Sort by any column to find largest/smallest/brightest objects
- Multiple row selection highlights multiple objects
This enables detailed inspection: “Which object has the largest area?” “What are the intensities of the brightest objects?”
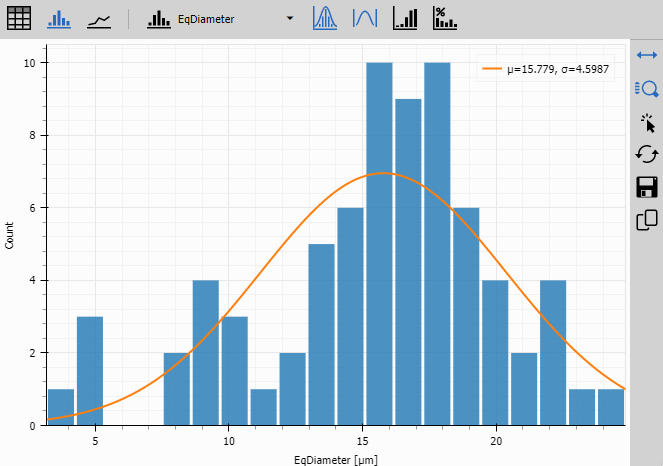
Histogram
Histogram shows the frequency distribution of any selected object feature.
Configuration:
- Data/Display: “Current frame” to show distribution for current frame objects
- Data/Displayed columns: Configurable feature selection (dropdown)
- Bin count: Number of histogram bins
- Fit normal: Optional Gaussian fit overlay
Visualization options:
- Select which feature to plot (Area, Circularity, Intensity, etc.)
- View distribution shape (normal, bimodal, skewed)
- Identify outliers or subpopulations
- Statistical fit shows mean and standard deviation
The histogram reveals patterns like:
- Uniform population (narrow distribution)
- Size heterogeneity (broad distribution)
- Two populations (bimodal distribution)
- Outliers (isolated bars far from the main peak)
Presentation block
The presentation block organizes all tables and graphs into a synchronized, interactive display. This creates a cohesive analysis interface where all views are linked together.
Layout structure
The presentation uses two Stacked Layout nodes to organize visualizations:
Left pane:
- Object Features Table (current frame individual objects)
Right pane (stacked vertically):
- Counts Table (all frames population statistics)
- Histogram (feature distribution)
- Line Chart (temporal trends)
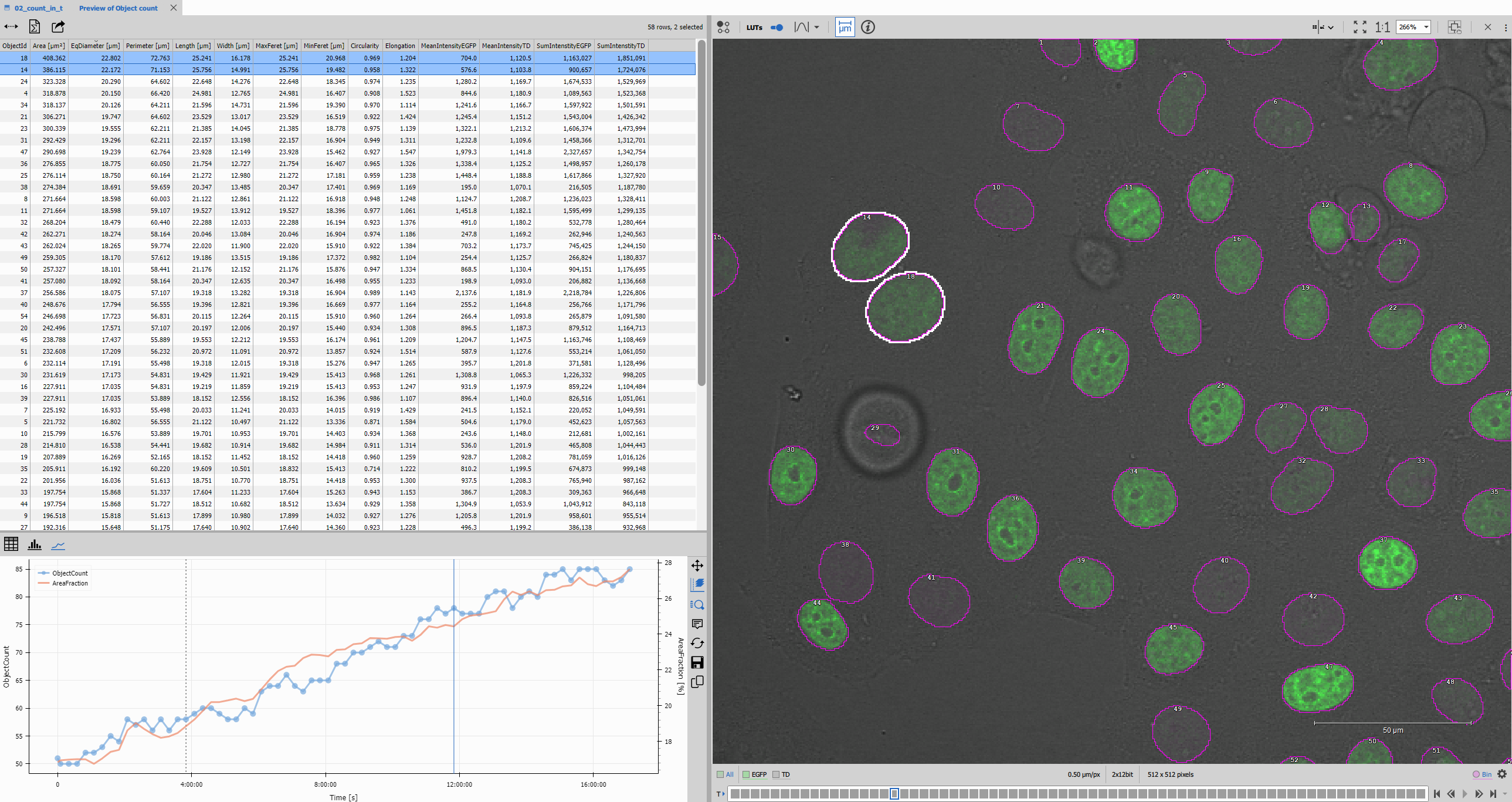
Display node
Display combines the two layout panes with the image viewer.
Settings:
- showNotes: 0 (hides the notes panel)
Synchronized interactions:
- Selecting an object in the image → highlights its row in Object Features Table
- Selecting a row in Object Features Table → highlights the object in the image
- Selecting a row in Counts Table → navigates to that frame
- All views update together when changing frames
The Display node creates a complete analysis environment where image data and measurements are tightly integrated.
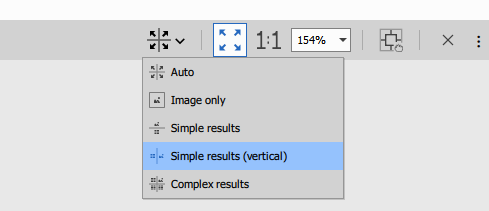
Layout customization
The layout button in the interface allows reorganizing views:
- Horizontal or vertical splits
- Resize panes by dragging dividers
- Show/hide individual views
- Maximize specific views
Experiment with layouts to find the arrangement that best suits your analysis workflow.
Output files
The recipe saves three types of output for further analysis or documentation:
Save Channels
Save Channels doesn’t modify channel data as it “stores” the source (Original):
- All input channels (RGB or fluorescence)
- Original intensity values preserved
Save Binaries
Save Binaries saves the segmentation results:
- Binary layers are saved into H5 file by default
- Useful for verifying segmentation quality
Save Tables
Save Tables saves all measurement results:
- Data is saved into H5 file
- Object count table (population statistics)
- Object features table (individual measurements)
- Can be exported to Excel, CSV for external analysis
- Results section includes all graphs and visualizations
Adding the Object Catalog (optional)
The catalog variation adds a visual gallery of all detected objects, which is especially useful for quality control and classification.
Additional nodes for catalog version
- Removes objects touching the image frame
- Prevents incomplete objects from appearing in the catalog
- Placed immediately after Threshold
- Creates a grid gallery of object thumbnails
- Input: Connected to the object features table
- Settings:
- tableRowVisibility: “all” to show all objects
- colorChannelSelection: Enabled for channel selection
- binaryLayerSelection: Enabled to show segmentation overlay
Modified layout:
- Object Catalog gets its own stacked layout pane
- Typically displayed on the left alongside the object table
- Creates a three-pane layout: Catalog | Table | Counts/Charts
Interactive features:
- Click a thumbnail to select that object in the image
- Thumbnails show the object cropped from the original image
- Can toggle between channels and binary overlay
- Useful for visual inspection of segmentation quality
The catalog is particularly valuable when:
- Verifying segmentation accuracy across many objects
- Identifying misclassified objects
- Finding objects with specific morphologies
- Quality control in automated screening
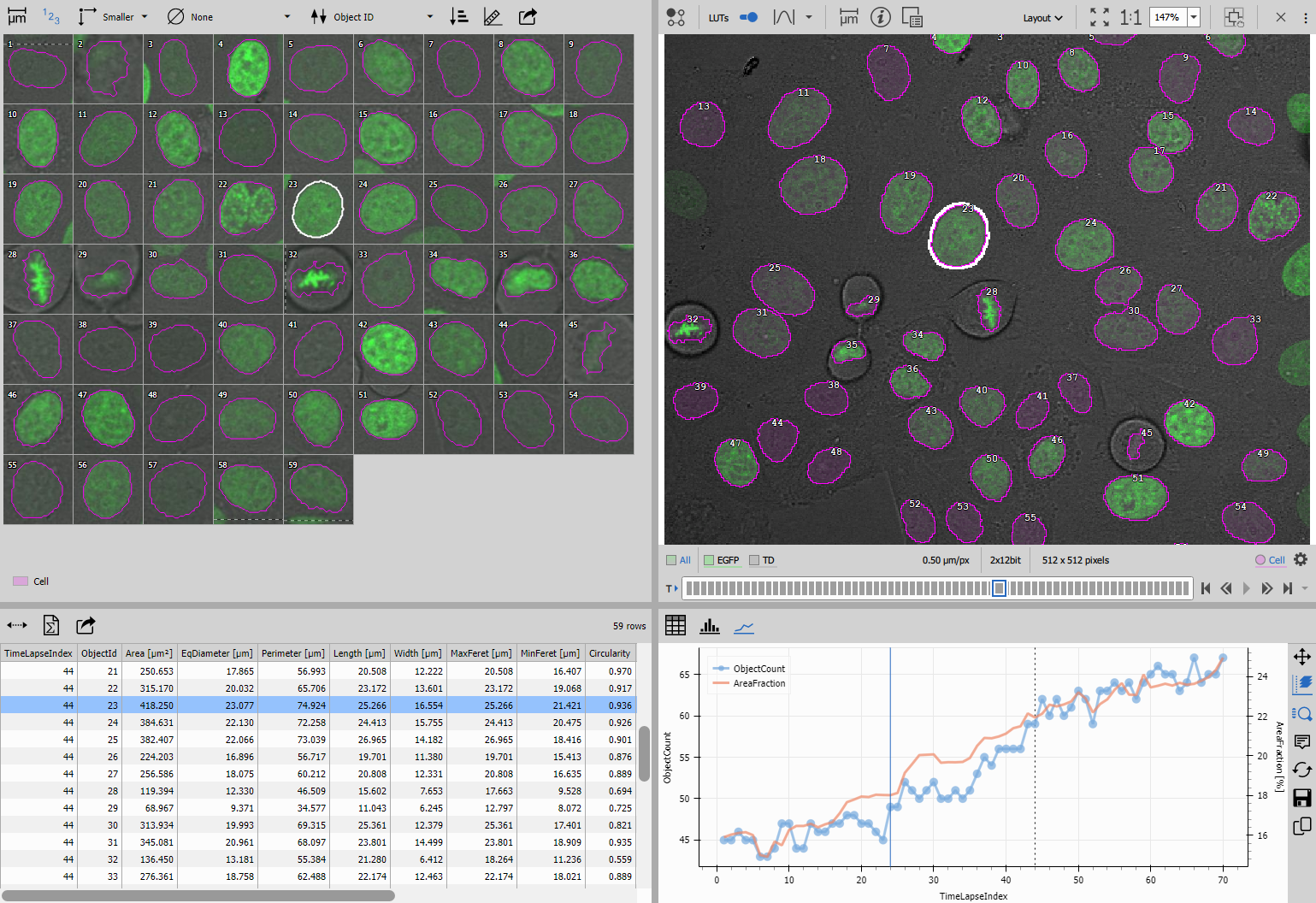
Extending the workflow
This basic counting workflow can be extended in many ways:
Add classification:
- Use feature thresholds to classify objects (small/large, dim/bright)
- Create calculated columns for classification logic
- Color-code objects by class
Add tracking:
- Connect objects across time using Track Objects
- Measure per-track features instead of per-frame
- See Tracking workflow
Add multi-well analysis:
- Process well-plate data with Wellplate Metadata
- Compare object counts across wells
- See Well-plate assays workflow
Add statistical analysis:
- Calculate p-values between conditions
- Fit curves to temporal data
- Use statistical nodes in Data manipulation category
Add 3D analysis:
- Replace 2D threshold with Threshold 3D
- Use Object 3D Count
- See 3D counting workflow
Related workflows
- Time measurement - Temporal measurements without individual object detection
- Cell measurement - Advanced cell morphology analysis with nucleus and cytoplasm
- Tracking - Connect objects across time
- Well-plate assays - High-throughput screening applications
- 3D counting - Volumetric object counting
Summary
Object counting is the foundational workflow in GA3 that demonstrates the complete analysis pipeline: segmentation → measurement → accumulation → visualization. While simple in concept (just threshold + count), this workflow teaches the essential patterns used in more complex analyses:
- Frame-by-frame vs. accumulated processing
- Current frame vs. all data display modes
- Field-level vs. object-level measurements
- Synchronized multi-view interfaces
Master this workflow and you’ll understand the building blocks for all GA3 analyses.