Deconvolution and AI restoration
Open the workflow in NIS-Express:
Deconvolution
The deconvolution package offers a suite of powerful tools aimed at restoration of clear images from acquired data degraded by distortions that take place in an optical microscope. The typical effects of such distortions include:
- Blurring of the sample structures around and below the resolution limit of the system
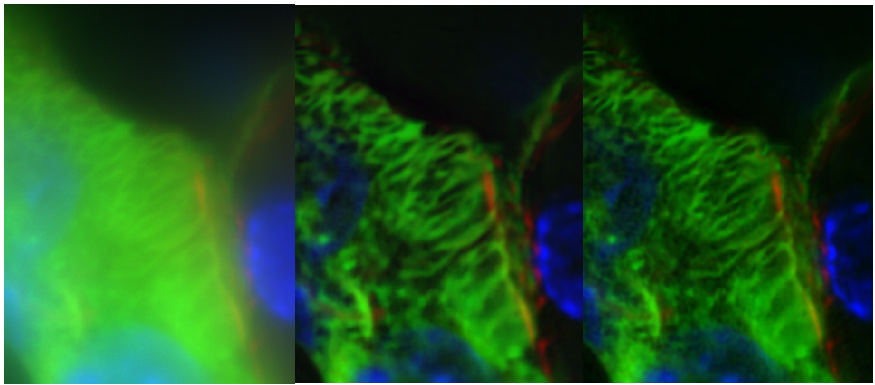
- Contamination of the focused Z plane with out of focus light
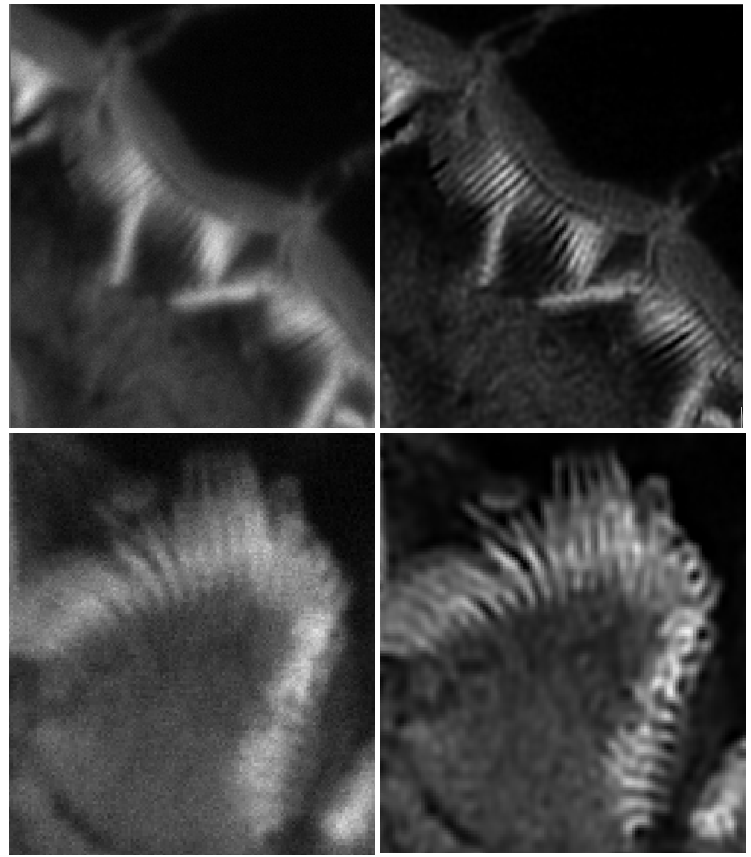
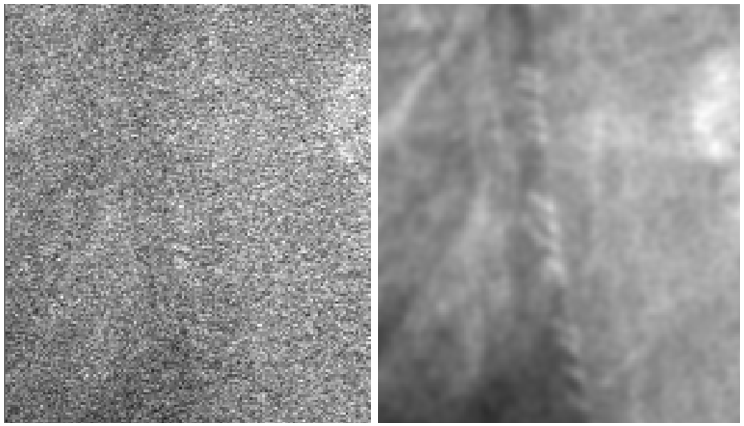
- Signal corruption by shot noise
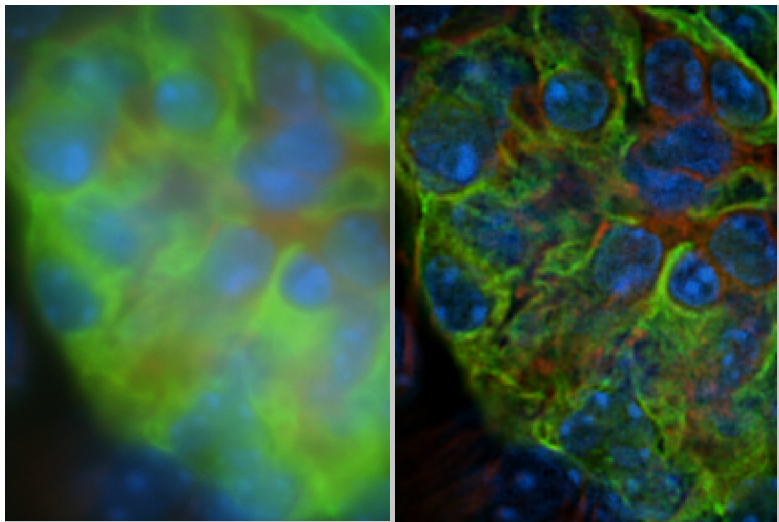
- Additional blurring induced by a mismatch between refractive indices of the immersion medium and the sample
Mathematically, the blurring operation can be modeled and expressed as a convolution operation. Hence the name deconvolution – the inverse of this process.
The deconvolution module provides several algorithms because there are multiple mathematical strategies to solve the problem, each with its own strengths and limitations. Since no single algorithm is optimal for all modalities, sample types, or imaging conditions, offering multiple options enables you to experiment and find the most suitable approach for your specific dataset.
The two most frequently used algorithms are MAP and Richardson-Lucy. For both, we also provide convenience variants that automatically estimate and set all parameters. These are called MAP Automatic and Automatic, respectively.
As a rule of thumb, we recommend starting with MAP Automatic or Automatic, and only proceeding to manual parameter tuning if the results are not satisfactory.
MAP (Maximum A Posteriori)
A state-of-the-art, versatile iterative algorithm suitable for all modalities. It offers mathematically proven convergence and is usually the best choice for wide-field and confocal data.
Richardson-Lucy
An iterative algorithm with fast convergence. It performs well on confocal, NSPARC, and spinning disc modalities, but may be slower and less effective for wide-field imaging.
Landweber
An iterative algorithm based on a Gaussian noise assumption.
Blind
An iterative algorithm which allows the PSF (i.e., the mathematical model of the microscope) to change during the deconvolution process. Less numerically stable, but can be a good choice for challenging, far-from-ideal imaging conditions.
Fast
A non-iterative, fast algorithm suitable only for high signal-to-noise ratio SNR datasets.
AI restoration tools
Standard deconvolution methods usually produce excellent results, but in some cases they may be too slow or not sufficiently effective at removing out-of-focus haze - especially for the wide-field modality.
If strict mathematical rigor or quantitativity is not required, our deep learning-based tools can often deliver very high-quality results, even for challenging samples.
Clarify.ai
A fast neural network designed for out-of-focus haze removal in wide-field images. For lower SNR datasets, it includes an option to preprocess the image using Denoise.ai.
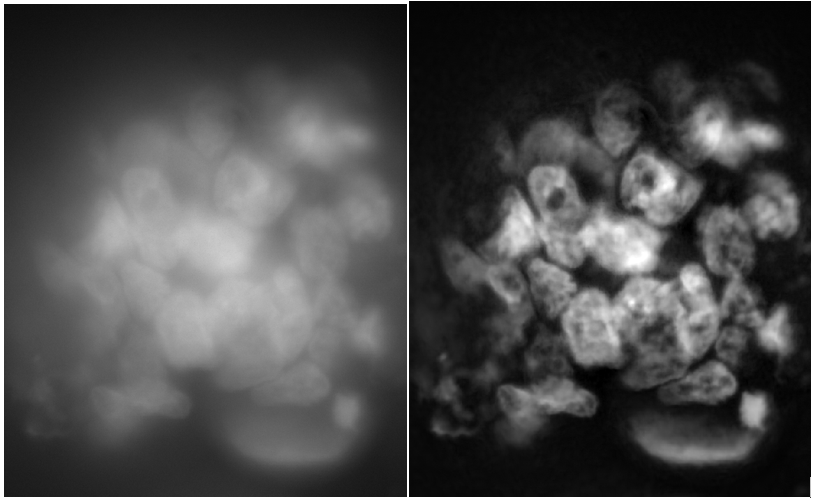
Restore.ai
A more complex AI-based solution that combines multiple end-to-end trained networks for denoising, deconvolution, and haze removal. Can be used with any imaging modality, but is generally slower than Clarify.ai.