Image processing
Image processing (sometimes called preprocessing) is used to process the image data (channels) for the analysis that follows (especially Segmentation and Measurement).
With novel segmentation methods (Segment.ai, SegmentObjects.ai, Cellpose) the need for traditional preprocessing like contrasts, convolutions, detection and morphological operation is diminishing. However, when the analysis requires absolute intensity values (Threshold, Growing to background) it is necessary to fix background unevenness or frame-to-frame intensity variations.
In many cases it is a practical strategy: distort the image to be easily thresholdable. For instance in case of a DIC image, cells can be segmented by detecting edges, smoothing it out and thresholding them.
Preprocessing to be always considered:
Deconvolution
Deconvolution and spectral unmixing are computational techniques that dramatically improve fluorescence microscopy images by correcting optical limitations and separating overlapping signals.
Deconvolution
Deconvolution removes blur and enhances resolution through mathematical modeling.
Deconvolution methods
- MAP (Maximum A Posteriori)
- A state-of-the-art, versatile iterative algorithm suitable for all modalities. It offers mathematically proven convergence and is usually the best choice for wide-field and confocal data.
- Richardson-Lucy
- An iterative algorithm with fast convergence. It performs well on confocal, NSPARC, and spinning disc modalities, but may be slower and less effective for wide-field imaging.
- Landweber
- An iterative algorithm based on a Gaussian noise assumption.
- Blind
- An iterative algorithm which allows the PSF (i.e., the mathematical model of the microscope) to change during the deconvolution process. Less numerically stable, but can be a good choice for challenging, far-from-ideal imaging conditions.
- Fast
- A non-iterative, fast algorithm suitable only for high signal-to-noise ratio SNR datasets.
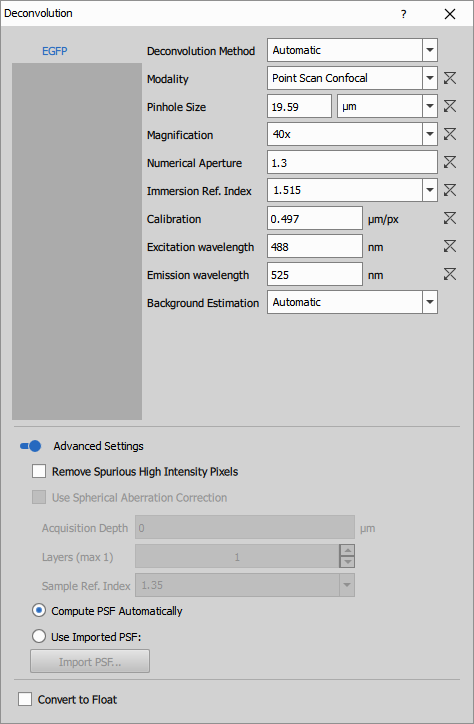
Parameters
See also: Deconvolution (group), Deconvolution workflow
Fluo. Unmixing
Remove fluorescence crosstalk and extract separated dye signals.
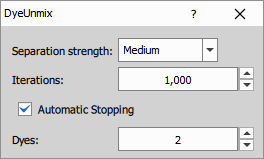
Parameters
See also: Deconvolution (group)
Background
Background signal in microscopy images originates from multiple sources including camera dark current offset, ambient light, sample autofluorescence, scattered excitation light, and optical artifacts.
More problematic is uneven background intensity across the field of view, caused by vignetting (light falloff toward image edges), uneven illumination from the light source, optical aberrations, or specimen-dependent effects like varying tissue thickness. These spatial variations create intensity gradients that interfere with segmentation, quantitative measurements, and image stitching. Background correction nodes remove these unwanted contributions to reveal true sample signal.
Auto Shading Corr.
Automatically detects and corrects vignetting and uneven illumination across the field of view. The algorithm analyzes the image to determine background type (bright, dark, average, or auto) and applies appropriate correction.
This method is particularly valuable before stitching multi-tile acquisitions, where uncorrected vignetting creates visible seams at tile boundaries. The correction works on each 2D plane independently in 3D datasets.
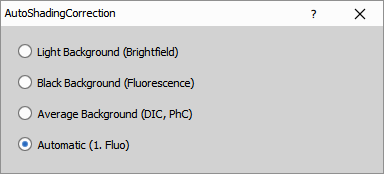
Select on which background the correction is performed:
- Light Background (Brightfield)
- Black Background (Fluorescence)
- Average Background (DIC, PhC)
- Automatic
Parameters
See also: Background (group)
Dark Offset (CMOS)
Newly scanned ND2 files captured with a CMOS camera may include metadata describing the camera’s dark offset. This node reads the offset value (when present) and automatically subtracts it from the input image.
Parameters
See also: Background (group)
Rolling Ball
Estimates and removes smooth background variations by rolling a sphere of defined radius over (bright backgrounds) or under (dark backgrounds) the image intensity surface. The radius parameter, specified in micrometers or pixels, should exceed the size of the largest object in the image—typically 1.5-2× the diameter of cells or other features of interest.
This method excels at correcting uneven illumination before segmentation, as it preserves local contrast while removing large-scale intensity gradients. Unlike global methods, rolling ball adapts to local background levels, making it robust for specimens with spatially varying background.
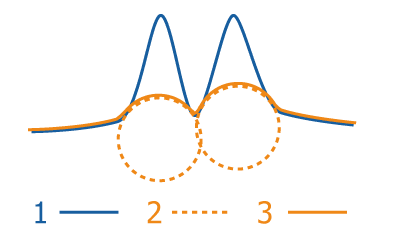
The image intensity is in blue (1), the rolling ball is in dashed orange (2) and the estimated background is orange (3). The final image is the distance between blue and orange.
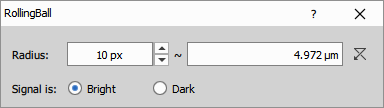
Parameters
See also: Background (group)
Subtract Background
Identifies the most frequent intensity value within a masked region and subtracts it from the entire image.
The mask defines either background regions (where the mode intensity is calculated) or object regions (where background is everything outside the mask). This approach works best when you can identify representative background areas, either interactively drawn or from previous segmentation steps.
The method provides simple, direct background removal but assumes uniform background intensity across the image.
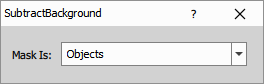
Parameters
See also: Background (group)
Subtract Constant
Subtracts a fixed intensity value from all pixels in the image.
This operation is essential for removing camera dark current offset before performing quantitative analyses like intensity ratios, correlation coefficients, or Manders’ overlap calculations, where a positive baseline offset would bias results.
Unlike adaptive methods, this provides reproducible correction but cannot handle spatially varying background.
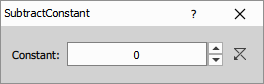
Parameters
See also: Background (group)
Ax Shading Correction
Removes shading artifacts specific to Nikon AX and AX R confocal microscopes.
This correction is tailored to the optical characteristics and illumination patterns of the AX line scanning confocal system. The node operates automatically without user parameters, applying manufacturer-optimized correction based on the microscope model metadata in the image file.
Parameters
See also: Background (group)
Contrast
Contrast nodes enhance image contrast by increasing the intensity difference between objects and the background. Low contrast images appear flat and gray, making segmentation difficult and obscuring subtle details.
Contrast adjustment is particularly important when intensity ranges vary between frames (as in time-lapse sequences where photobleaching occurs), when objects have similar brightness to background, or when uneven illumination creates local variations that global thresholding cannot handle. The nodes in this group offer both global adjustments that operate on the entire image histogram and local methods that enhance contrast adaptively within neighborhoods.
Auto Contrast
Automatically stretches image contrast by mapping a percentile range of pixel intensities to the full output range.
The Low parameter (e.g., 0.01) defines the bottom percentile that becomes black, while High (e.g., 0.01) defines the top percentile clipped to white, with the remaining 99.98% of pixels linearly stretched between these extremes.
This method normalizes intensity variations between frames in time-lapse sequences, ensuring consistent brightness for threshold-based segmentation that relies on absolute intensity values. The automatic percentile-based approach is more robust than min-max stretching, as it ignores outlier pixels from noise or hot pixels.
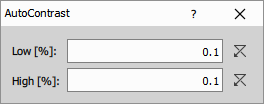
Parameters
See also: Contrast (group)
Contrast
Provides manual control over contrast adjustment using multiple transfer functions:
- Linear (standard min-max stretching),
- Logarithmic (compresses bright values, expands dark),
- Exponential (opposite of log),
- Equalized (global histogram equalization redistributing intensities uniformly), and
- Gamma correction (power-law transformation).
The Low and High parameters define input intensity values that map to minimum and maximum output, while Gamma controls the curve shape for power-law transforms. This node offers maximum flexibility for tailoring contrast to specific needs—logarithmic for high dynamic range, gamma for midtone adjustment, equalization for revealing detail across the entire intensity range—making it suitable when auto methods fail or specific visualization requirements exist.
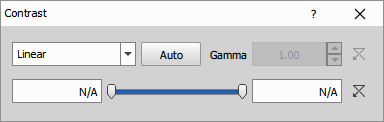
Parameters
Input
- A (Channel): Input image to be processed
Output
- R (Channel): Output processed image.
Control
Low (Number): A value that will become the minimum intensity (0).
High (Number): A value that will become the maximum intensity.
Type (Number): Linear, Logarithmic, Exponential, Equalized, Gamma correction
Gamma (Number): Gamma correction (where applicable).
See also: Contrast (group)
Gauss-Laplace
Enhances edges and fine detail by applying a Laplacian of Gaussian (LoG) filter, implementing unsharp masking to boost high-frequency components.
The Power parameter (range 1.01 to 2) controls enhancement strength by interpolating between the original image (Power=1, no enhancement) and maximum sharpening (Power=2, full LoG contribution).
Unlike simple sharpening, the Gaussian smoothing component reduces noise amplification while the Laplacian detects edges, making this method more sophisticated than raw Laplacian filtering. This approach enhances object boundaries and internal texture without over-emphasizing noise, useful before segmentation of low-contrast objects or for improving visual quality of final images.
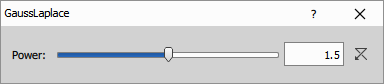
Parameters
See also: Contrast (group)
Local Contrast
Enhances contrast by adjusting pixel intensities relative to their local neighborhood rather than the global image histogram, making it ideal for images with uneven backgrounds or spatially varying illumination.
The Radius parameter defines the neighborhood size for local analysis, while Degree controls the strength of contrast amplification, with higher values producing stronger local enhancement. This adaptive approach reveals objects that would be invisible with global methods—for example, dim cells in one region and bright cells elsewhere can both be enhanced appropriately.
The method is particularly valuable before segmentation when rolling ball background subtraction is insufficient or when objects have locally variable contrast against background.

Parameters
See also: Contrast (group)
Sharpen
Increase sharpness by applying a Laplacian filter.
The node doesn’t have a control dialog.
Parameters
See also: Contrast (group)
Sharpen Slightly
Increase sharpness by applying a mild Laplacian filter.
The node doesn’t have a control dialog.
Parameters
See also: Contrast (group)
Ultra Details
Enhance ultra details (Alpha, Details, Noise).

Parameters
See also: Contrast (group)
Unsharp Mask
Increase sharpness by subtracting a blurred version of the image.
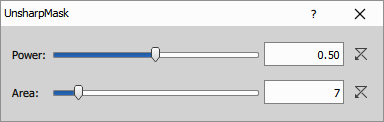
Parameters
Input
- A (Channel): Input image to be processed
Output
- R (Channel): Output processed image.
Control
Power (Number): Set the strength of the sharpening effect. The possible range of values is from 0.01 to 1.
Area (Number): Define the area size which is used during calculating the result. The larger the area is, the smoother the result will be.
See also: Contrast (group)
Convolution
Convolution is a fundamental mathematical operation that applies a small matrix (kernel) to each pixel in an image, computing weighted sums of neighboring pixels to produce filtered output. The kernel slides across the image, and at each position, pixels within the kernel’s footprint are multiplied by corresponding kernel weights and summed to determine the new pixel value.
Different kernels produce different effects: averaging kernels smooth images by blurring, derivative kernels detect edges, and Gaussian kernels provide controlled smoothing that preserves important features better than simple averaging.
Convolution forms the basis for many image processing operations including smoothing, sharpening, edge detection, and noise reduction. For more details, see Convolution on Wikipedia and Kernel below.
Common Filters
Applies predefined 3×3 convolution kernels selected from a list of standard types.
The small 3×3 size makes this operation fast but limits smoothing power and edge detection scale—only immediate neighbors influence each pixel. This node is useful for quick preprocessing like mild smoothing before segmentation or edge enhancement, but larger-scale smoothing requires Gaussian filters with adjustable radius.
Filters:
- Sobel (east, north, south, west): Directional edge detection (see Wikipedia)
- Prewittt (east, north, south, west): Directional edge detection (see Wikipedia)
- Laplacian (4, 8): Edge detection
- Sharpen more: Aggressive sharpening
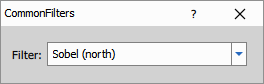
Parameters
See also: Convolution (group)
Gaussian Filters
Choose from Gaussian and Gaussian derivative filters to modify the image.
Filters:
- Gaussian, (X and Y): Blurs the image.
- Gaussian ∂x, Gaussian ∂y: First derivative. Detects (emphasizes) edges in the specified axis direction.
- Gaussian ∂x2, Gaussian ∂y2: Second derivative. Detects (emphasizes) edges in the specified axis direction.
Parameter σ (dispersion) for the Gaussian filter. For the basic Gaussian filter, it determines extent of the blur. For the other filters, it determines the size of edges that are going to be detected.
Parameters
See also: Convolution (group)
Golay Filter
Use Golay filter to smooth the image or enhance edge detection.
Also known as Savitzky–Golay filter (see wikipedia).
Can be used to
- Smooth the image and
- Detect edges
Parameters
See also: Convolution (group)
Mexican Hat
Use Mexican Hat filter to improve edge detection or fine image details.
The Mexican Hat kernel is defined as a combination of a Laplacian kernel and a Gaussian kernel. It enhances edges and also reduces some noise (see wikipedia).

Parameters
See also: Convolution (group)
Denoising
For the state-of-the-art AI denoising use the Denoise.ai.
Below are conventional imaging methods for Denoising and Smoothing.
Denoising
Reduce noise by using a stack of frames as reference.
Denoising is used to reduce any kind of noise in the image (Gaussian, Poisson noise). The described neighborhood is 3D – information is taken from the neighboring frames.
Denoising method:
Original method is based on denoising via large neighborhoods in a spatial and frequency (incl. wavelets coefficients) domain. It is a one-pass algorithm.
Regression algorithm iteratively denoises every pixel according to its local neighborhood. In every iteration, local linear regression is computed for every pixel and the value is replaced by the regressed value. This method belongs to the same family as the Original method.
Fusion between Original and Regression. It takes the better parts of both algorithms.
Bayes uses a probabilistic approach is used to compute the most likely estimate from the neighboring pixels.
Iterative prediction is very similar to the Regression method, but in each iteration a new value is determined by voting based on extrapolations from neighboring pixels. Note that this method is slow.
Denoising Power defines the strength of the denoising algorithm for each channel separately. If it is set to zero, denoising is done with an estimated noise variance. Move the slider to the right and denoising calculates with higher noise variance (more noise is present in the image) and vice versa.
Parameters
See also: Denoising (group)
Denoising (frame)
Reduce noise by using a single frame as reference.
Same as the Denoising except it works on 2D neighborhood and is much faster.
Parameters
See also: Denoising (group)
Destriping
Remove stripes created by VT-iSIM device.

Parameters
See also: Denoising (group)
Low Pass Filter
Remove high-frequency components and smooth the image.

Parameters
See also: Denoising (group)
Median
Minimize noise by replacing pixels with the median value of their neighbors.
Parameters
See also: Denoising (group)
Smooth
Smooth the image with a Gaussian filter.
Parameters
See also: Denoising (group)
Smooth (fast)
Quickly smooth the image with a Gaussian filter approximation.

Parameters
See also: Denoising (group)
Detection
DIC Objects
Enhance contrast of DIC objects for easier thresholding.
The differential interference contrast imaging method produces a special type of image in which objects appear three-dimensional, with a bright edge on one side and a shadow on the other. Such objects cannot be thresholded and further analyzed. The Detect DIC Objects command performs recognition of the objects and makes them bright in order to be easily thresholded.

Parameters
Edges
Highlight pixels at sharp intensity changes, revealing edge boundaries.
The node doesn’t have a control dialog.
Parameters
Filaments
Highlight neurons, blood vessels or similar structures.
Detects and highlights 2D/3D filament structures (e.g. neurons, blood vessels, …) present in the image. Set the diameter range of the filaments in your sample, set how many times this range is evaluated (Steps) and enhance the result using Contrast.
Parameters
Gabor Edge Detect
Highlight edges based on direction and strength.
Gabor Edge Detection (see wikipedia) method represents a linear filter used for visualizing specific features of an image. Use the Orientation combo box to select a direction [°] in which the features are visualized. Select a proper direction and adjust the Amplitude of the wavelet to find the sharpest result.
Real, Imaginary or Both filters can be applied. Switching from Positive to Negative value highlights the surroundings of the object.
- Orientation: Select a direction in degrees in which the features are visualized.
- Amplitude: Adjust the amplitude of the wavelet to find the sharpest result.
- Filter Apply one of the filters or both.
- Use Value: Switching from Positive to Negative highlights the surroundings of the object.
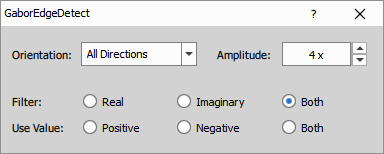
Parameters
Input
- A (Channel): Input image to be processed
Output
- R (Channel): Output processed image.
Control
Orientation (Number): The direction in degrees in which the features are visualized.
Amplitude (Number): The Amplitude of the wavelet to find the sharpest result.
Filter (Number): Real, Imaginary or Both.
Value (Number): Positive, Negative or Both. Positive to Negative highlights the surroundings of the object.
Gradient Morpho
Highlight edges using the difference between dilation and erosion.
Parameters
See also: Morphology (group)
Peaks
Highlight local maxima.
Parameters
See also: Morphology (group)
Regional Maxima
Highlight regional maxima using top-hat transformation.
Parameters
See also: Morphology (group)
Regional Minima
Highlight regional minima using top-hat transformation.
Parameters
See also: Morphology (group)
Valleys
Highlight local minima.
Parameters
See also: Morphology (group)
Insert
Gradient Image
Create a smooth gradient with defined brightness stops.
Inserts a grayscale color gradient over the connected image. The gradient is defined by a gradient table where the first column sets the color change points and the second column defines the color of the segment (0 = black, 1 = white). Use decimals to set a specific change position or shade of gray. Angle of the gradient can be adjusted optionally.
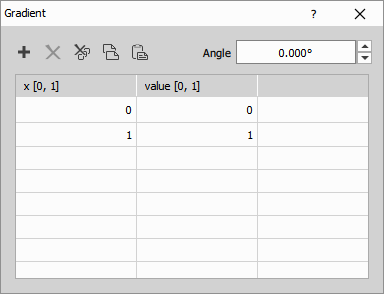
Parameters
Morphology
In image processing, a structuring element matrix, also known as a kernel or a filter, is a small rectangular or square grid which acts as a template that slides over the image, and at each position, it interacts with the neighboring pixels to produce a new value for the pixel in the output image. The size of the matrix, also referred to as the kernel size, determines the extent of the neighborhood considered for the computation.
The element is ran more than once to effectively increase its size.
Different square matrix shapes available:
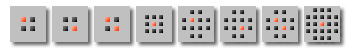

Some functions support a “circular” matrix where different distances from the central pixel are compensated.

- Matrix: Click the button to change the structuring element used for this operation.
- Count: Number of iterations: effectively the size of the operation.
Open
Erode to remove small bright areas and dilate to restore outline
Parameters
See also: Morphology (group)
Close
Dilate to remove small dark areas and erode to restore outline
Parameters
See also: Morphology (group)
Erode
Expand dark regions
Parameters
See also: Morphology (group)
Dilate
Expand bright regions
Parameters
See also: Morphology (group)
Fill Holes
Fill enclosed dark areas inside bright regions with surrounding intensity
Parameters
See also: Morphology (group)
Linear Morphology
Linear morphology is similar to morphology except it uses a special set of linear Matrices as structuring elements.

See also: Morphology
Linear Open
Erode along a line to remove small bright areas and dilate to restore outline
Parameters
See also: Linear Morphology (group)
Linear Close
Dilate along a line to remove small dark areas and erode to restore outline
Parameters
See also: Linear Morphology (group)
Linear Erode
Erode along a line to expand dark regions
Parameters
See also: Linear Morphology (group)
Linear Dilate
Dilate along a line to expand bright regions
Parameters
See also: Linear Morphology (group)
Labels & Classes
Smooth
Smooth pixel variations and create patches.
Parameters
Transformations
Add Borders
Extend image borders.
Adds grayscale color borders to the connected image. Enter the border size [px] for each side (Top, Left, Right, Bottom) and enter the color value (0=black, MAXVAL=white).
Parameters
Input
- A (Channel): Input image to be processed
Output
- R (Channel): Output processed image.
Control
Left (Number): Border size in pixels.
Top (Number): Border size in pixels.
Right (Number): Border size in pixels.
Bottom (Number): Border size in pixels.
Value (Number): Intensity value to be filled into the border pixels.
Binning
Merge neighboring pixels into bins and reduce image size.
- Binning factor of NxN. For example 3x3 will reduce the area of 9 pixels into a single pixel in the resulting image based on the selected method.
- Binning method The resulting pixel value will be calculated as maximum, minimum, mean or sum. Sum No Noise method brightens the image but does not amplify noise like the sum method does.
Parameters
Input
- A (Channel): Input image to be processed
Output
- R (Channel): Output processed image.
Control
Factor (Number): Binning factor of 3x3 will reduce the area of 9 pixels into a single pixel in the resulting image based on the selected method.
Method (Number): Resulting pixel value will be calculated as maximum, minimum, mean or sum. Sum No Noise method brightens the image but does not amplify noise like the sum method does.
Crop
Crop the image to the given rectangle.
Parameters
Change Canvas
Change the canvas dimensions, centering the original data without resizing.
Parameters
Flip
Flip the image horizontally or vertically.
Parameters
Move
Move the image horizontally and vertically.
Parameters
Resize to Calibration
Scale the image to match the calibration.
Parameters
See also: Resize in ND processing & conversions
Resize to Ref
Resize the image to match the size of the reference.
Parameters
See also: Resize in ND processing & conversions
Rotate
Rotate the image around its center.
Parameters
See also: Rotate in ND processing & conversions
Lin. Morphology 3D
Linear morphology 3D is similar to linear morphology. It operates on 3D volumes and in the Z direction. The structuring element is
[1 1 1] over Z.
See also: Linear Morphology and Morphology
Linear Open Z
Erode along a Z-axis to remove small bright areas and dilate to restore outline
Parameters
See also: Lin. Morphology 3D (group)
Linear Close Z
Dilate along a Z-axis to remove small dark areas and erode to restore outline
Parameters
See also: Lin. Morphology 3D (group)
Linear Erode Z
Erode along a Z-axis to expand dark regions
Parameters
See also: Lin. Morphology 3D (group)
Linear Dilate Z
Dilate along a Z-axis to expand bright regions
Parameters
See also: Lin. Morphology 3D (group)