Installation
NIS-Express installation is available in the following two variants:
- Basic:
NIS-Express-Basic-X.YY.ZZ.exeor - Advanced:
NIS-Express-Advanced-X.YY.ZZ.exe
where the X.YY.ZZ is the version number.
The installer performs a non-administrative install for the Current user only. The program is installed into Programs (by default) under the current user Home folder.
Program Files for all users as Administrator use the /ALLUSERS switch:
NIS-Express-Advanced-X.YY.ZZ.exe /ALLUSERSTo reveal the destination folder
To reveal the destination folder paste the path into the navigation bar in the Windows File Explorer.
Current user
%userprofile%\AppData\Local\Programs
All users
C:\Program Files\
Optional components
Basic variant
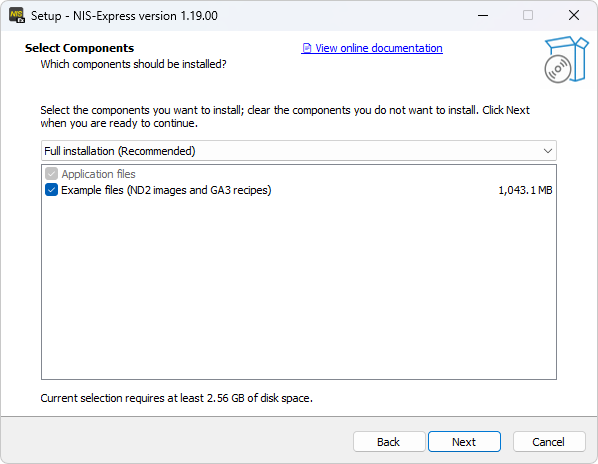
Advanced variant
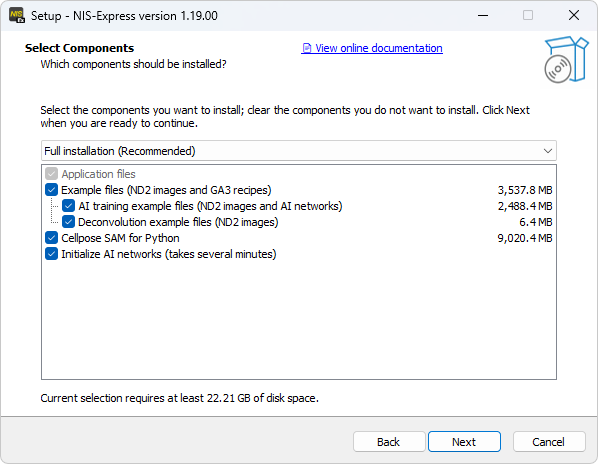
Example files with Credits
Example ND2 images and GA3 recipes is a good starting point for users not familiar with the GA3 or NIS-Elements.
| Image files | GA3 recipe files or workflow |
|---|---|
| 01_cell_count.nd2 | 01_object_count_RGB.ga3 01_object_catalog_RGB.ga3 |
| 02_count_in_t.nd2 | 02_object_count_fluo.ga3 02_object_catalog_fluo.ga3 |
| 03_time_meas.nd2 | 03_time_meas_channels.ga3 03_time_meas_rois.ga3 |
| 04_cell_meas.nd2 | 04_cell_meas.ga3 |
| 05_meas_moving.nd2(1) | 05_meas_moving objects.ga3 |
| 06_tracking.nd2 | 06_tracking.ga3 |
| 07_spt.nd2 | 07_single_particle_tracking.ga3 |
| 08_live_dead.nd2 | 08_live_dead.ga3 |
| 09_tracking_3d.nd2 | 09_tracking_3D.ga3 |
| 10_counting_3d.nd2(1) | 10_cell_meas_3D.ga3 |
| 11_align.nd2 | ND > Align Frames |
| 12_edf.nd2 | ND > EDF |
| 13_root_mono.nd2 | ND > Difference of subsequent frames |
| 14_stitch_mp.nd2 | ND > Stitch multi-point into large image |
(1) Nikky Corthout, VIB BioImaging Core Leuven, BE
Examples will be installed in users’ Documents directory in NIS-Express folder.
Image files:
%userprofile%\Documents\NIS-Express\Images\ExamplesGA3 recipe files:
%userprofile%\Documents\NIS-Express\Ga3Recipes\ExamplesAI training example files
Example ND2 images and AI networks are useful too see how to train each network.
| AI networks | File for training and validation |
|---|---|
| Segment.ai | 21_segment_obj_train.nd2 21_segment_obj_validation.nd2 |
| Segment Objects.ai | 22_segment_nuclei_train.nd2 22_segment_nuclei_validation.nd2 |
| Enhance.ai | 23_enhance_train.nd2 23_enhance_validation.nd2 |
| Convert.ai | 24_convert_hela_train.nd2 24_convert_hela_validation.nd2 |
Examples will be installed in users’ Documents directory in NIS-Express folder.
Image files:
%userprofile%\Documents\NIS-Express\Images\ExamplesAI network files:
%userprofile%\Documents\NIS-Express\AiNetworks\ExamplesDeconvolution example files
| Image files | Workflow |
|---|---|
| 31_decon_2d.nd2 | Deconv > Deconvolution Deconv > Clarify.ai |
Examples will be installed in users’ Documents directory in NIS-Express folder.
Image files:
%userprofile%\Documents\NIS-Express\Images\ExamplesPython Cellpose 4 SAM for GA3
Installs Cellpose Segment anything model (SAM) environment into PythonEnvs folder:
%userprofile%\AppData\Roaming\Laboratory Imaging\NIS-ExpressInitialize AI networks
Initializes and stores the networks in a form suitable for the current GPU. If unchecked the initialization will be done before first use of each network.