Recipe page
The Recipe page is were the GA3 recipes are created, modified and run.
This is a typical workflow on this page:
Open image file
Open files using Image button.
Create a recipe
Open one of the predefined workflows from the toolbar, create a new from scratch or open an existing recipes from the recipe list.
Edit the recipe
Edit the recipe nodes and its parameters in the GA3 editor while observing the changes in the Main window.
Run it
Finally Run the recipe.
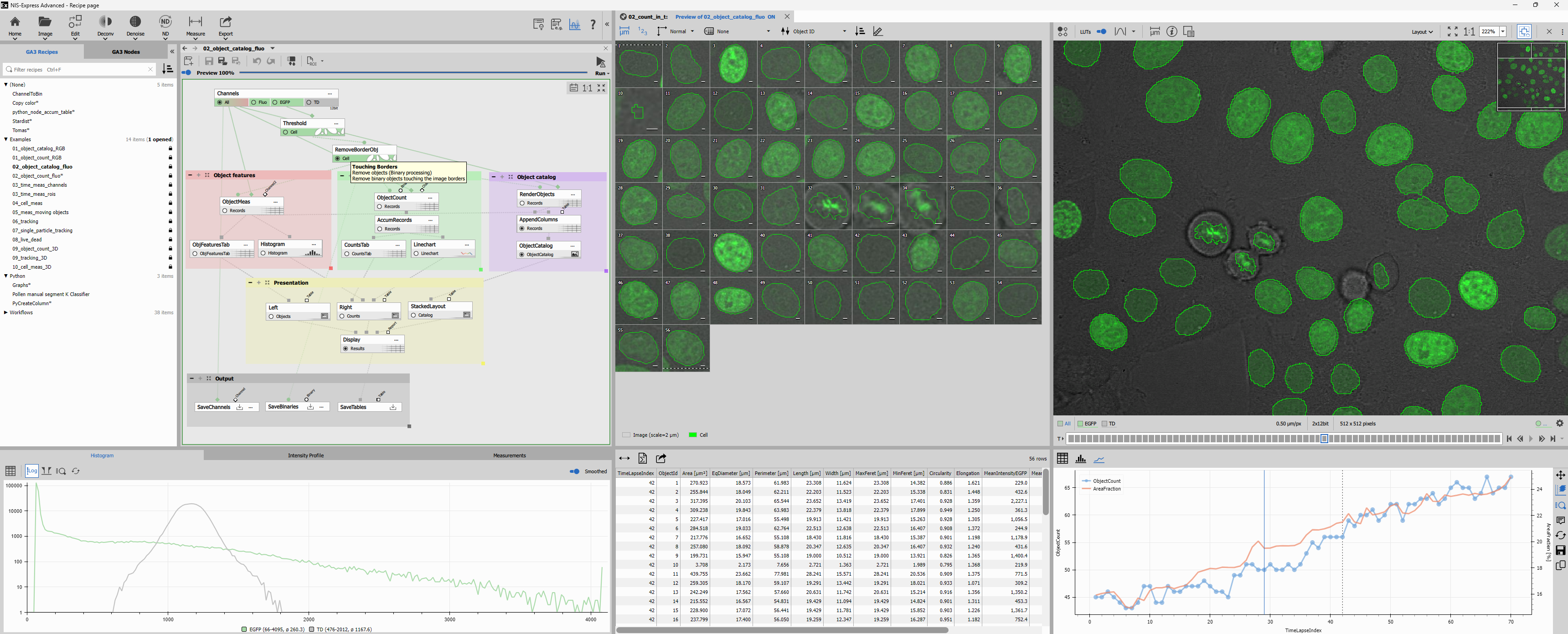
Main toolbar

Returns back to the home screen. To go to specific page use the dropdown:
- Image page Alt+I - opens the current image in the Image page.
- AI page Alt+A - opens the current image in the AI page.
Image button shows Open image dialog. The bottom down arrow contains more options:
- Recent images - lists recently opened images for direct opening.
- Recent folders - lists recently visited folders for opening a file from that particular folder.
- Paste image - creates a new image from the clipboard (it is disabled when the clipboard doesn’t contain a bitmap or image).
- Import file… - opens single-file conversion. See Import single image.
- Convert images… - opens batch conversion for individual files. See Import individual files.
- Import image sequence… - opens sequence conversion. See Import image sequence.
Open all related results from the document ⋮ menu button in the top-right.The Edit operations are basic image modification tools. Except the Crop image they are GA3 recipes in a wizard. When the the recipe is Run a new document is created. On Cancel the the wizard reverts back to the underlying recipe.
- Crop image - shows a dedicated dialog where the crop can be defined. Specifically:
- a rectangle in the image,
- ND dimensions (frames in Z-stack, time-lapse or multi-point),
- channels and
- binary layers
- Resize - shows the Resize node dialog where new size can be specified in pixels or percent.
- Rotate - shows the Rotate node dialog where the angle and cropping mode can be defined.
- Flip horizontally / vertically - flips the image using the Flip node.
- Change bit-depth - shows the Change Bit Depth node dialog where the desired bit-depth can be specified.
- Convert to RGB - converts the image using the Render Color node into a new image file.
- Convert to 8-bit RGB - converts the image using the RenderColor node and ChangeBitDepth node into a new image file.
- Convert to Mono - converts the image using the Channel to Intensity node into a new image file.
- Convert to 8-bit Mono - converts the image using the ChannelToIntensity node and ChangeBitDepth node into a new image file.
- Import labeled binaries - shows the Labels to Binary node dialog where a filename or a sequence of binary labeled files can be specified.
The deconvolution package offers a suite of powerful tools aimed at restoration of clear images from acquired data degraded by distortions that take place in an optical microscope.
– see the Deconvolution workflow
- Deconvolution - shows the Deconvolution node dialog.
- Clarify.ai - shows the Clarify.ai node dialog.
- Restore.ai - shows the Restore.ai node dialog.
Denoising removes unwanted corruption of acquired images by noise, which is inherently present in the image formation process of a microscope.
– see the Denoise workflow
- Denoising - shows the Denoising node dialog.
- Denoise.ai - shows the Denoise.ai node dialog.
ND processing function operate on a whole dimension (sequence of frames). For more details and image examples see the ND processing workflow.
- Align frames - aligns images in a Z-Stack or a Time-lapse using Align node.
- Extended depth od Focus (EDF) - makes focussed image from a Z-Stack using EDF node.
- Maximum intensity projection (MaxIP) - creates a maximum Intensity projection (brightest value over a loop for every pixel) using Max IP node.
- Average of all frames - calculates an average of values over a loop for every pixel using Average node.
- Median of all frames - calculates a median of values over a loop for every pixel using Median node.
- Difference of subsequent frames - calculates a difference between values of subsequent frames over a loop for every pixel using Select Frame node and Subtract node.
- Stitch multi-point to large image - merges multiple images from different XY position into a single large image using Stitch Multi Points node dialog.
Measure functions contain ready-to-use recipes which produce image analysis results like tables, graphs.
Object count - segments objects and produces (see the workflow):
- table with object counts per frame,
- table with measured object features,
- histogram of all measured features per frame and
- time chart with the number of object and area fraction versus time
Object catalog - segments objects and produces all as the Object count plus (see the workflow):
- interactive Object catalog
Time measurement (channels) - measures channels intensity (optionally in a ROI) and produces (see the workflow):
- table with field intensity features and
- time chart of measured intensities versus time
Time measurement (ROIs) - measures channels intensity (on multiple ROIs) and produces (see the workflow):
- table with field intensity features in every ROI and
- time chart of measured intensities in Every ROI versus time
Measure cells - measures cell-specific features and produces (see the workflow):
- table with cell, cytoplasm and nucleus features and
- scatterplot of any feature versus another one
Measure moving objects - measures object features on cells that are moving (have to be tracked) and produces (see the workflow):
- table with object measurement of all objects in all frames organized by object/track,
- time chart of any object feature versus time and
- visualization of objects/tracks in the image
Tracking motility - tracks objects, measures motility features on moving objects and produces (see the workflow):
- table with object measurement of all objects in all frames organized by object/track,
- table with motility measurements per track and classification (steady, motile, progressive),
- time chart of any object feature versus time and
- visualization of objects/tracks in the image
Single particle tracking - tracks spots, measures position, calculates MSDs and produces (see the workflow):
- table with particles per tracks,
- table of MSDs per track,
- plot of single aggregated MSD and
- visualization of objects/tracks in the image
Live-dead assay - detects all cells and dead cells, counts both and produces (see the workflow):
- table with well dosing and live, dead, all counts and Z`,
- bar chart of live/dead per frame,
- wellplate schema with thumbnails, dosing, heatmaps and bar chart per well and
- dose response curve a Live ratio vs. dose
3D object count - segments 3D objects, counts them and produces (see the workflow):
- table with object counts per volume,
- table with measured object features,
- histogram of all measured features per frame and
- time chart with the number of object and area fraction versus time
3D tracking - tracks 3D objects, measures motility features on moving objects (hence have to be tracked) and produces (see the workflow):
- table with object measurement of all objects in all volumes organized by object/track,
- table with motility measurements per track,
- scatterplot of motility any feature vs a different one,
- chart of the relative object movement of all objects and
- visualization of objects/tracks in the image
3D cell measurement - segments 3D cells, measures and counts them and produces (see the workflow):
- table with object counts per volume,
- table with measured object features and
- histogram of all measured features per frame
Export function can take snapshots (RGB screenshots), presentations (snapshots of image and results) and export raw data into TIFFs and OME-TIFFs,
Snapshot to clipboard takes a snapshot of the image as seen.
Save snapshot… saves a snapshot of the image as seen.
Save full-res snapshot… saves a snapshot of the image as seen in full image resolution.
Save movie… saves a snapshot of the image as seen in full image resolution into a movie.
Presentation to clipboard takes a snapshot of the whole document area into the clipboard.
Save presentation… saves a snapshot of the whole document area.
Save presentation movie… saves a snapshot of the whole document area into a movie.
Save image as TIFF… saves the current frame into a TIFF.
Dataset as OME-TIFF… saves the ND image into OME-TIFF.
Dataset as series of TIFFs… saves the ND image into a series of single plane TIFFs.
The buttons at the right toggle the:
- Get started panel
- Examples panel
- Interactive tools panel and
- link to this help page
List of recipes
Available list of GA3 recipes is shown on the left alternatively with GA3 nodes palette.
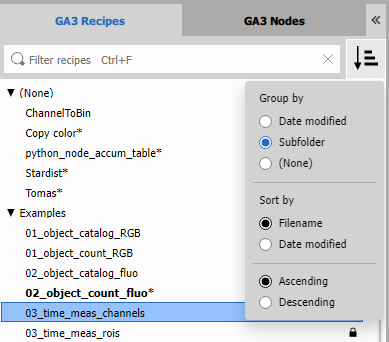
All GA3 files from a predefined folder (see settings) are listed. The list is synchronized with the corresponding folder – any change to the folder is reflected in the list and vice versa. Recipes can be organized in single-level sub folders (here “Examples”). Recipe opened in the editor is shown in bold such as 02_object_count_fluo.
Modified recipes are marked with an asterisk (*). Their modified state is saved in a filename with a tilde ~ appended
alongside the original file.
Viewing
List viewing options are accessible in the drop-down menu .
- Grouping
- Recipes can be grouped by:
- date when the file was last modified or
- subfolder one level below the selected folder.
- Sorting
- The recipes can be shown in ascending or descending order based on their:
- name or
- date.
Actions on recipes
Following actions are available on selected recipe(s) or the current one:
- Open Enter: opens the current recipe in the editor
- Insert into current recipe: inserts the current recipe (snippet) into the editor
- Marking as favorite: marks current recipe with favorite icon and adds the Recipe to the Run menu on Image page
- Show run history…: opens a list of ND2 (original) files which the current recipe was run on
- Save/Revert changes…: saves the modified version of the current recipe or discards it
- Copy Ctrl+C: copies the url of the current recipe to clipboard
- Copy recipe name: copies the name of the current recipe to clipboard
- Copy recipe path: copies the full-path and filename of the current recipe to clipboard
- Rename… F2: renames the current folder or recipe
- Delete… Del: deletes the current folder or selected recipes
- New Folder… : creates a new folder
- Refresh F5: refreshes the list of recipes
- Open in File Explorer… Ctrl+E: opens the recipe folder in the File Explorer
Recipes can be dragged out (only current recipe) into different applications. And GA3 files can be dragged in from windows explorer.
When grouped by folders Drag and Drop is available to move recipes between folders.
Filtering
To show only a subset of recipes start typing into the search field and the view will update accordingly.
Rules & examples
- Basic search
- Simply type words to find recipes whose filename contains them.
object countfinds recipe with both object and count in the filename.
- Exact phrases
- Use
"quotes “…” to search for an exact phrase.
"object count"finds recipe with both object count in the filename.
- Excluding words
- Add a
-minus sign before a word or phrase to exclude it.
object -countfinds recipe with object but not count.
- Tags
- Use
#tagto search for recipes with that tag or shortcut.
#modfinds modified recipes.
| tag | meaning |
|---|---|
#ro | read-only |
#mod | modified |
Modify-time related shortcuts:
#today, #yesterday, #thisweek, #lastweek, #thismonth, #lastmonth, #thisyear, #lastyear
- Dates
- Search by modify date
mtimefields with comparison operators.
Format:
YYYY-MM-DD(full date)MM-DD(month and day only)
Operators: <, <=, =, >=, >, !=
mtime:<2024-05-10finds recipes modified before May 10, 2024.
- Sub-folders
- Search by sub-folder name.
sub:examples
s:examplesfinds all recipes in a sub-folder that contains examples in the name.
- Combining searches
- Use
ORfor alternatives.
object OR tracking- Grouping
- Use parentheses
(…)to group terms:
(object OR tracking) 3Dfinds recipes containing 3D and either object or tracking.
- Wildcards
- Use templates with wildcards
- * matches any number of characters.
- ? matches exactly one character.
recipe*matches recipe, recipe2024, etc.
- Tip
- You can mix and match these rules — for example:
#ro date:>=2024-01-01 "object count" -draft*finds recipes in 2024 mentioning object count, excluding any draft files.
GA3 Editor
The editor consists of:
- Node palette: list of all available nodes
- Editor toolbar: manages the preview and the currently open recipe
- Editor main view: canvas where the nodes are connected into the recipe
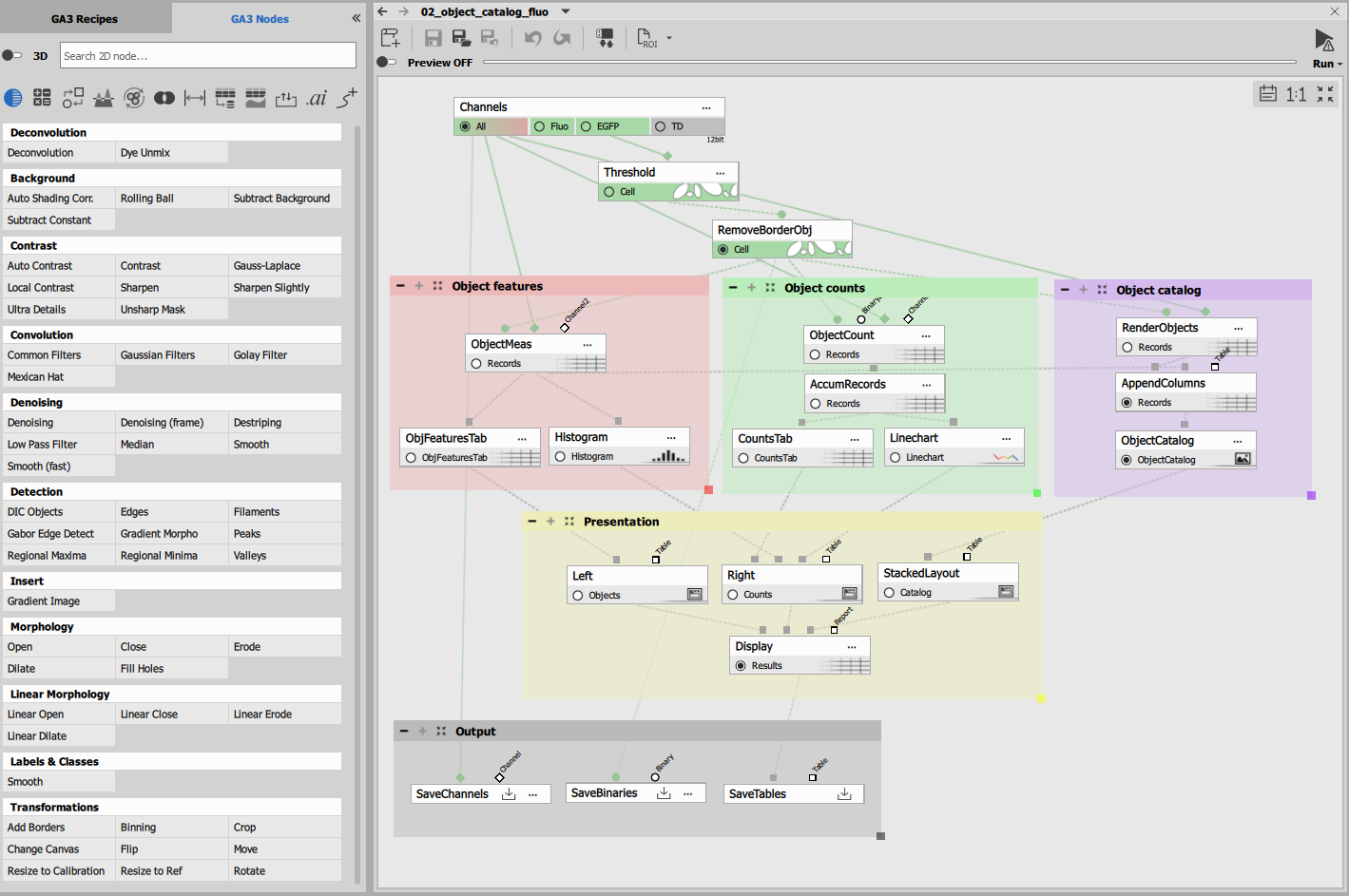
Recipes
Recipes can be:
- opened from
- the list for recipes,
- the File explorer by dragging them inside the editor or
- created from
- the toolbar edit and workflow buttons,
- the New or Import buttons in the editor toolbar.
- Auto-saving
- The GA3 editor never calls for saving a recipe. Instead it silently saves the recipe (alongside the original as recipe~.ga3).
- Unsaved recipes
- When a new recipe is created it has an
[unsaved]tag near its name until they are explicitly saved. ‘Unsaved’ recipes are stored in a temporary folder for a limited time until newer unsaved recipes push it away. - History
- Recently opened recipes (including unsaved) can be accessed in the recipe history using arrow back or a dropdown menu near the recipe name.
- Attached document
- A GA3 recipe is always attached to one document where its preview is displayed. The attached document’s name displays a pin icon. When another document is activated the recipe is not reattached to it automatically. Instead a pin icon is shown in the editor to let the user attach it manually.
Snippets
Are ordinary GA3 recipes (usually organized under snippets subfolder) containing relatively small functionality stored
for later use as a building block for more complex recipes. As being normal recipes, snippets are easy to share.
Use “Insert into current recipe” context menu item to use the snippet. The Channel Input and Save Output nodes are automatically stripped and new nodes are inserted as a new section.
Ideal for:
- specific segmentation, processing and operations
- python nodes
Node palette
The palette lists all GA3 nodes organized by their function into:
- Categories – each on different page and
- Groups – separating the nodes on a page.
The 3D switch exchanges the node variants between 2D and 3D where applicable.
First row of icons represents node categories. The order follows the order in which they are typically used in a recipe:
- Starting with operations on image channels (image preprocessing, operations between images, ND & conversion).
- Segmentation for detecting objects of interest called binaries or binary layers.
- Operations on binaries (binary processing, binary operations).
- Measurement nodes that produce data tables with measured features on fields and objects.
- Data management to manipulate and analyze the data
- Results and Graphs for data visualization.
- Finally exports and imports and specialized nodes in AI and Tracking.
Nodes in the palette have typically a tooltip which further describes the function it performs. On right-click it shows a context menu with a link to the reference in this help.
To insert the node into the recipe:
- drag&drop it in the editor,
- click on the node to place it below the last node
- double-click it to auto-connect it to the last node
Nodes can be searched for using
search field.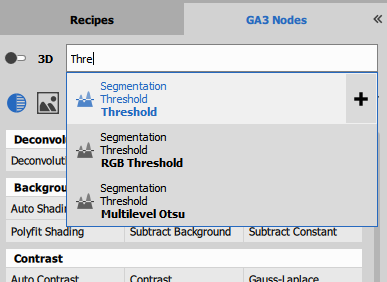
Editor toolbar
- New recipe
- Clears the current recipe.
Remember: to go back use the arrow back near the “New recipe [draft]”. - Save
- Save the current recipe into the original file (deletes the recipe~.ga3)
- Save As
- Save the current recipe into new recipe file.
- Revert back the last saved state
- Loads the original recipe.ga3 (deletes the recipe~.ga3).
- Undo
- Goes back in the current recipe undo stack.
- Redo
- Goes forward in the current recipe undo stack.
- Import & export
- Imports and exports current recipe from a ga3 file and exports HTML and PNG files.
- ROI
- Defines a smaller rectangle and a subset of frames to speed up the analysis preview.
- Preview accumulate records
- Switches on/off the “Apply only on current frame in preview” for all the accumulated records. Speeds up the preview.
- Attach document (only on different document)
- When different document is selected it must be attached manually in order to start preview on it.
Remember: currently attached file has its name in accent color. - Preview
- The switch enables the preview. If enabled any change to the recipe will trigger update.
- Run
- Runs the analysis on the whole file (on all frames).
Editing and preview
Editing the recipe consists of connecting suitable nodes into a graph and setting their parameters in a way that it produces the desired result.
To mark a node’s output for preview check the appropriate radio button. Note that consecutive outputs with the same name will check exclusively. In the following recipe the preview shows the “RGB” channels, the “Bin” binary layer and “Records” dataset table.
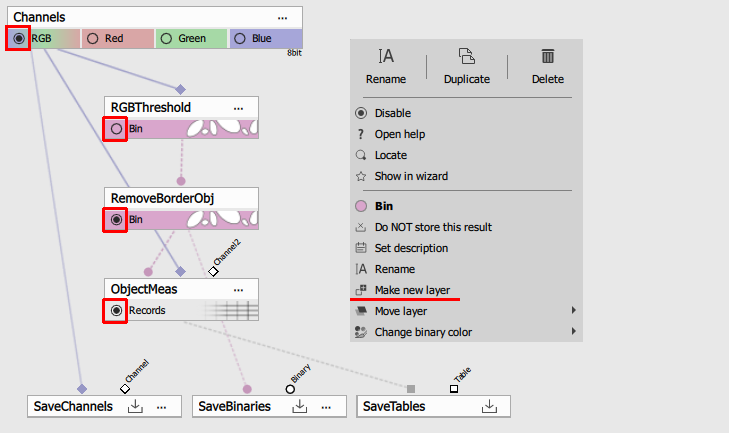
If the “Remove Border Object” node’s Bin output is check for preview the same “Bin” output of the “RGB Threshold” is unchecked. In order to see both the down-stream Bin should be a different binary layer. To do so select the “Make new layer” from the context menu and check both radio buttons.
Note that what is displayed in preview is different from what is connected to the Save nodes and actually saved when the recipe is run.
The preview updates to reflect the changes made to the recipe. The results of the intermediate nodes are cached and therefore not calculated over again. Only the edited node and all down-stream nodes are recalculated. However, when modifying a node which is upwards in the graph it may take a while to process the dependent nodes. Similarly, nodes that have to go over many frames (ND processing, Tracking, …) to produce their output take longer time.
Therefore, to improve preview speed use preview ROI to crop the image or/and the number of frames to reduce the amount of data to be processed while maintaining useful preview. Alternatively, when doing many edits at once it may be faster to switch OFF the preview entirely.
Remember that the preview doesn’t necessarily process all frames or all nodes. It does only what is necessary to show the results of nodes marked for preview.
Running the recipe
The run can be set either to:
- overwrite previous results (e.g. when running the same recipe with different settings) or
- to create a new result (e.g. to compare different results of different parameters in a single recipe).
Files
When a recipe is run it creates or modifies files (ND2, H5) based on what the recipe does. Specifically what is connected into the Save* sink nodes.
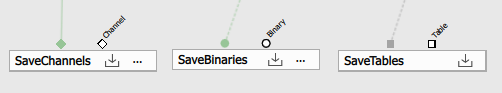
Do not confuse what is shown in the preview with what is actually saved when running the recipe!
When a recipe (e.g. Decon.ga3) modifies image channels (a.k.a. color data) on an existing image
(e.g. image.nd2) a new ND2 file is created. In this case it will be named image - Decon.nd2.
Any other output (binary, tables, results, …) of a recipe is stored inside an HDF5 file alongside the original ND2 with the same file base name as the ND2. The H5 file can contain more than one analysis result.
When running a CellCount.ga3 recipe that saves only binaries and tables and does not modify the
image channels on the image.nd2 file it will produce a new H5 file image.h5.
When image channels are modified and other outputs are saved by the recipe a new ND2 file and a new
H5 file is generated. In this case it will result in image - DenoisedRatio.nd2
and image - DenoisedRatio.h5.
This logic preserves the original ND2 files.
- Decon.ga3
- Decon10it.ga3
- Decon20it.ga3
- DenoisedRatio.ga3
- image.nd2
- image.h5
- image - Decon.nd2
- image - Decon10it.nd2
- image - Decon20it.nd2
- image - DenoisedRatio.nd2
- image - DenoisedRatio.h5
Documents
An Image in Recipe page is called:
- Image Original: until changed by the preview
- Image Preview of Recipe: after the preview
- Image Recipe: after the recipe was run
- Image | Recipe | Another_recipe: after another recipe was run
When more recipes were run the results may be compared by selecting between the Recipe and Another_recipe. They can be compared side by side too.
Wizard
In GA3, nodes can be marked as to be shown in the wizard. If done so, a special GUI with only marked dialogs are presented to the user.
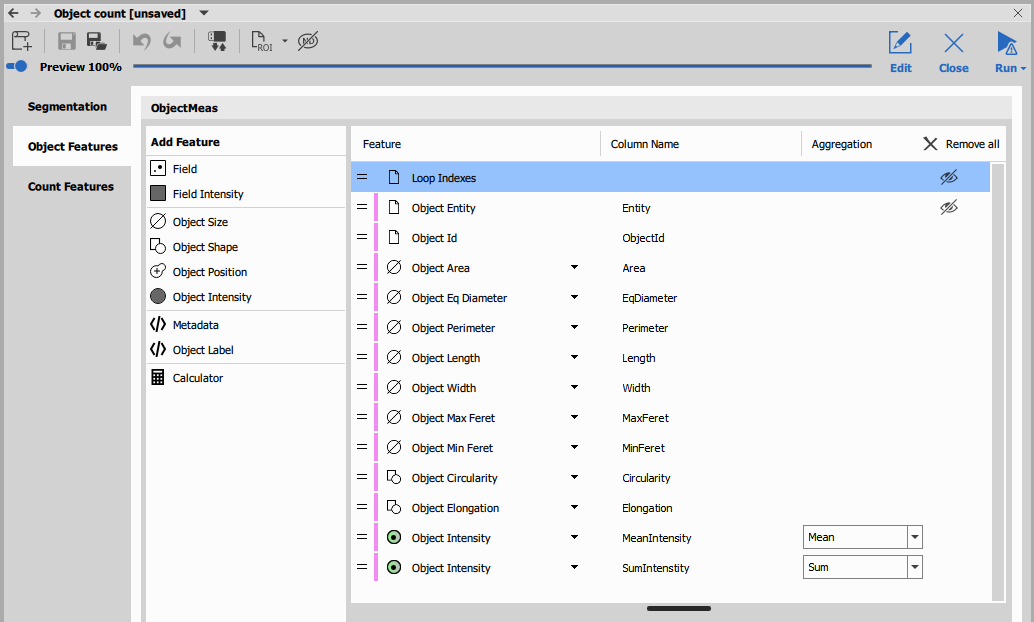
Nodes’ GUIs can be grouped into pages and special nodes can be used to show specific preview on each page (see the wizard nodes).