General Analysis 3
Overview
GA3 is a software tool for performing image processing and analysis. It does so by executing a recipe which defines what has to be done specifically step-by-step. Individual processing steps are called nodes. The nodes are organized into a graph that defines the execution flow.
Every recipe has one or more source nodes (e.g. Channels, Binaries, …) and some sink nodes (e.g. Save Channels, Save Binaries, Save Tables, …) acting as the recipe input and output.
Nodes have Inputs and Outputs that are of three types: Channel, Binary and Table. There are also Control parameters which are entered in the settings Dialog of the node and which are typically not part of the input and output. The (oriented acyclic) graph is built by connecting one nodes’ output to the next node input.
The node’s outputs have a Name and Color (tables have only name) to distinguish them one from the other. Most nodes typically only modify input/output hence they do not change these properties. However, some nodes (such as Segmentation, Measurement nodes) create new outputs (having a new name and color). Outputs can be renamed (and their color changed) making them a new entity. This is useful when a before and after output is needed for preview or to be stored.
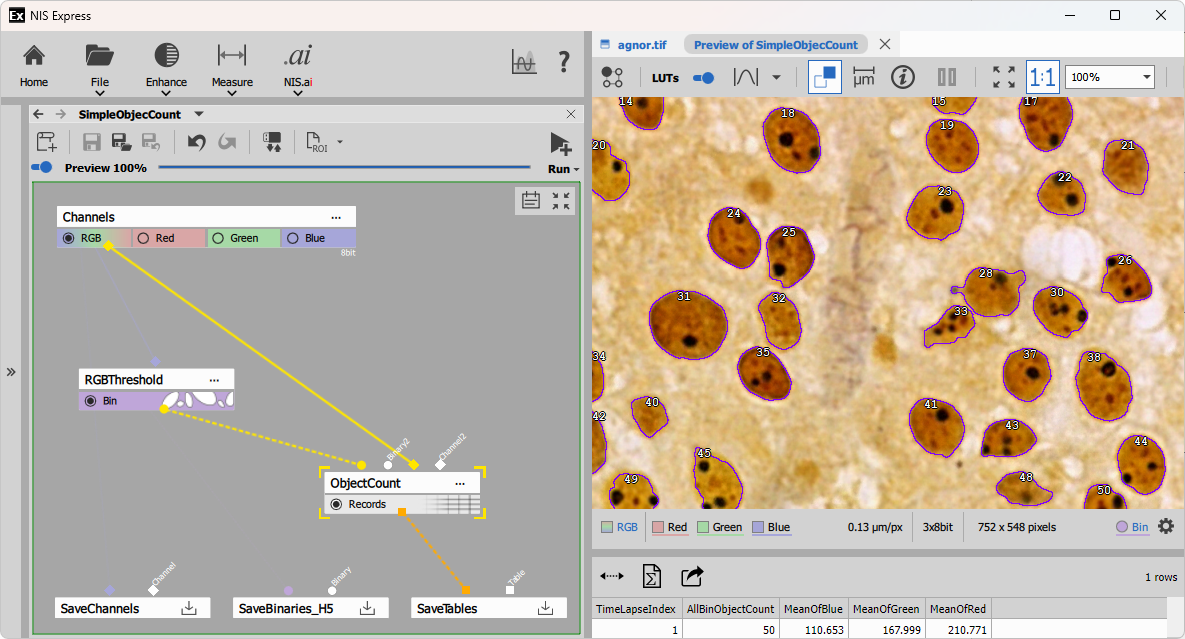
The example shows a simple object count recipe composed of two nodes:
- RGB Threshold that segments 50 objects outlined in violet and
- Object Count that counts the objects and measures Mean intensities in all objects shown as table with one row.
Nodes
There are hundreds of nodes that can be used in a recipe. However, there are only few that are used regularly. Others are used sparingly because of their niche role. The nodes are organized into categories and groups in the order as they are typically used.
Finally, the scientific question and image data dictate which nodes should be used and how complex the recipe needs to be.
Some workflows do only image processing (denoising, deconvolution, EDF, …) where the input is an image with some defects and the output is a better image.
However, typically the workflow is more complex and involves more steps:
Preprocessing
Image processing typically aims at improving the image properties for further analysis. Especially Segmentation. Most common image quality issues and nodes that solve them:
Excessive noise is a result of experiment needs with regards to the amount of time available, light dose the sample can endure or similar compromises. Use Denoise.ai to remove the noise or Denoising, Median, Smooth to reduce it.
Uneven field intensity comes from optical system shading/vignetting or sample features like auto-fluorescence. Use Auto Shading Corr. or Rolling ball to mitigate it.
Crosstalk between channels comes from light bleed-through optical filters. Use Fluo. Unmixing to reassign it back to the correct channels.
Sub-optimal resolution and confocality are caused by imperfections of the optical system. Use Deconvolution to improve confocality and aberrations by reassigning the light.
Segmentation
It aims at detecting segments of pixels in the image such as particles, cells, organoids, animals or just foreground in order to be further measured, counted and analyzed. AI based segmentation methods outperform the conventional methods. However there is tradeoff between time and quality/robustness:
Threshold is the simplest conventional method that is very fast works very well when the image quality is good: uniform background, good signal to noise ratio and pronounced objects. As it is pixel based method objects tend to split or merge incorrectly when the quality requirements are not met. For example Nuclei are typically successfully segmented on DAPI or Hoechst when image quality permits. Otherwise use Cellpose.
Bright spot detection is conventional method that excels at detecting circular spots of similar size that that peek above the noise (bright spots). It works very well in both 2D and 3D. For small spots always prefer Spot detection. Works surprisingly well with big spots too (a.g. animals). For transparent samples use Dark spot detection
Cellpose SAM is a popular pre-trained AI based community model that excels at segmenting cells including cytoplasm both in 2D and 3D. It works with H&E staining as well as with fluorescence. On the other hand it is much slower than conventional methods. Stardist is another popular community model falling into this category.
Segment.ai and Segment Objects.ai is an AI based method that excels where all previous methods fail. As it has to be trained it is very robust in segmenting the objects.
Binary postprocessing
The goal of postprocessing is to fix issues with the segmented objects so that they better represent the underlying structures. Here are the common ones (also part of the threshold node):
Too many objects with rough borders are caused by thresholding noisy or uneven image. Use Smooth to merge more little debris into meaningful objects.
Connected objects are caused by not being pronounced enough. Use Separate Objects to cut where objects are touching.
Too many small debris caused by noisy image. Use Clean to remove it.
Objects have holes due to their morphology. Use Fill holes or Close Holes to fill partial holes.
Unwanted objects are included because the segmentation algorithm cannot discriminate them. Use Filter Objects to select objects by its Size, Shape or Intensity. Remove objects touching border not to bias the measurements with incomplete objects.
Measurement
These nodes produce the quantitative data by measuring pixel/voxel intensity values, object counts, their size, shape and position and associate them with appropriate entities: fields, objects, cell etc.
Field, Volume measurements produce intensity (incl. ratiometric) features of 2D frames or 3D volumes. The measurements are typically performed during time or under different condition. Typical examples are Calcium, FRET, FRAP experiments.
Object measurements produce 2D or 3D object features such as: size, shape, intensity and position of spots, nuclei, cells, animals.
Object Count produces 2D or 3D object counts and aggregated object feature measurements (mean, min, max, standard deviation, etc.) for all objects on a frame or volume. The prime use of this node is counting one or multiple binaries – classes of objects.
Children and Parent measures two class of 2D or 3D objects: Children and Parents where one is inside of the other or in it’s zone of influence. The first measures children object features and aggregates with respect to parent objects and the second measures parent object features and aggregates of its children.
Cell produces measurements per 2d cells. It measures all the object features for the whole cell, nucleus and some on cytoplasm. It can quantify optional spots in all these compartments.
Data manipulation
Data manipulation nodes typically operate on measurements outputs – tabular data. The most common data manipulation is to bring data coming from different measurements to a single table.
Accumulate Records is a node that gets rows from all frames into single table. These accumulated data can be viewed, plotted into graphs or fitted. For memory and performance reasons the GA3 engine processes the multidimensional datasets frame-by-frame.
Append Columns and Join Records are nodes that combine measured data into single table. Whereas the former is simpler as it assumes same number and order of rows the later allows to specify how the rows should be matched. Where more than row corresponds to one row 1:N the one row is duplicated N times.
Reduce Records calculates statistics for each group of rows that is defined by one or more columns having a same value. For example a table is first accumulated and reduced by well, a drug concentration or a well label.
Presentation
The aim of presentation nodes is to visualize the data in tables or graphs like histograms or scatter-plots. It is also possible to display multiple panes to show more tables and graphs simultaneously and in a synchronized way using one of layout nodes.
Node parameters
Image Channels
Image data (sometimes called color or picture) are organized into channels. Channel names (and their number) are taken from the source file and dictated by the underlying experiment. Fluorescence images have channels like DAPI, FITC and so on whereas brightfield images are typically RGB such as H&E stained images.
The source Channels node provides special channel called All or RGB in case of RGB images. RGB and All are treated the same by most nodes. There are, however some nodes that require specifically RGB image. Processing nodes by default process all channels the same. Deconvolution node is the notable exception which allows defining per-channel settings.
The channels parameters have metadata which are assigned before the run of GA3 (during the design of the recipe).
Notably:
- bit-depth: 8, 10, 12, 14, 16 bits unsigned integer or 32 bit float image
- calibration: microns per pixel
- size: width, height
All channels must have same metadata when saved to the ND2 file. In case of difference they are coerced to their maximum value in the save node.
Binaries
Binaries (a.k.a Binary layers) are typically result of segmentation. They are made of contiguous patches or segments of pixels representing entities such as cells, spots or organoids. The are displayed as semi-transparent overlays or contours on the image.
The same principles apply to 3D where objects are made of contiguous 3D patches of voxels.
Binary objects may have an ID which links them to measured features stored in tables. In such case the binary is stored as an ‘image’ of 32 bit integers, where every pixel is a natural number (object ID) or zero (background).
In many situations the object ID is not needed and as its expensive to compute it pixels are simply marked as objects or background. Stored as 8 bit unsigned integer where zero is background and non-zero is object. These objects have implicit IDs going from top-left to bottom-right from 1 to N.
An object to be must not touch others (to be separate objects). It must be divided by one pixel horizontally and vertically.
Binaries have same limitations as Channels regarding metadata during GA3 run.
Tables
Tables are 2D arrays of values organized into columns and rows. They are primary result of measurement.
Columns in GA3 are typically made of one or more book-keeping (called system) columns such as loopIndex for every loop (time, z-stack, multi-point), object ID and entity for objects followed by feature columns.
Book-keeping columns are used to link a table row to frame, volume or object. If a table doesn’t have such columns it looses capability to link to the image data.
Columns have metadata such as:
- ID: column identifier
- Title: what is displayed in the column header
- Unit: displayed in
[a.u.] - Type: such as Number or text
- Display: number of decimal digits and notation
Book-keeping columns have always the same ID. Other columns have their column ID kept from the moment of their creation. The benefit being that changing the title doesn’t break the downstream nodes (such as graphs) which reference the columns by ID and update with the title change.
When merging tables however, columns may be discarded because of their ID which is same coming from two or more tables where only the leftmost is taken. In such situation use New Column ID node.
Rows are filled during the GA3 run. By default tables are processed frame-by-frame or volume-by-volume unless accumulated.
Rows values may contain various data encoded into text such as
- formulae as a result of fits,
- jpeg or png images as thumbnails.
Control parameters
Nodes are setup using settings dialogs. Each node have a different more less complex depending of its functionality. Some nodes do not have settings dialog at all.
The set of Control parameters represent the state of the node.
The widgets in the dialog correspond to Control parameters which are:
- Number
- Text
- Table
Dynamic parameters
With a special dialog some Control parameters can turn into input parameters making them dynamic.
This is useful when a control parameter (typically a number) has to be set by a preceding node. As an example threshold may take the low limit from minimum intensity or a quantile.
Multiple dimensions
GA3 handles multiple dimensions (ND) like Z-Stack, Time-lapse or Multi-point implicitly. By default it processes the input ND image frame-by-frame in the same order as the frames are stored in the file. The nodes in the graph may change this behavior. For example 3D nodes switch the processing to volume-by-volume.
Some nodes like Maximum Intensity Projection (MaxIP) or Accumulate records may reduce or completely collapse a dimension. In that case the following nodes operate only on the reduced dimension.
When collapsed outputs are connected into a node together with non-reduced output the former is silently broadcasted back to have same number of frames as the latter.
For example: An Average node when applied to the time-lapse will produce only one frame – image that contains only static parts. This single averaged image can then be subtracted from all frames. The Subtract node will treat the single frame coming from the Average node as if there were as many frames as in the whole time-lapse and do the subtraction on all frames (this is called broadcasting).
Building the GA3 graph – Recipe
Here there are some commonly used GA3 graph building patterns.
Image processing
The Image processing graphs typically connect one (or more) processing nodes one after the other from
the source Channels node through All channels and save the result.
Three examples below show three common processing nodes:
--- config: look: handDrawn theme: neutral --- graph TD in1[Channels] -->|All| p1["Deconvolution"]:::red -->|All| out1[Save Channels] in2[Channels] -->|All| p2["Denoise.ai"]:::red -->|All| out2[Save Channels] in3[Channels] -->|All| p3["Fluo. Unmixing"]:::red -->|All| out3[Save Channels] classDef red stroke:#d88
See also Deconvolution, Denoise workflows.
Stack reductions
Typical example of stack reduction for fluorescence images is Maximum intensity projection (MaxIP) as it brings the data (high pixel values) from all frames into a single image.
MaxIP can reveal
- trajectories in a time-lapse or
- object XY footprint in a z-stack.
--- config: look: handDrawn theme: neutral --- graph TD in1[Channels] -->|All| p1["MaxIP"]:::red -->|All| out[Save Channels] classDef red stroke:#d88
In many cases it is practical to calculate a projection or Reduction over a loop and then subtract it from every frame.
In the examples below:
the output of Median contains static part of scene which can be Subtracted from each frame. This is helpful for segmentation when detecting moving objects.
variance over time-lapse is not built in, but can be constructed (as any higher moment) using builtin nodes using the formula: where can be replaced with Average node.
Subtract previous frame from the current with Select Frame.
---
config:
look: handDrawn
theme: neutral
---
graph TD
in1[Channels]
in1 -->|All_N| sub1
in1 -->|All_N| p1[Median]:::red -->|All_1| sub1[Subtract]:::red -->|All_N| out1[Save Channels]
in2[Channels] -->|All_N| sub2
in2 -->|All_N| p2[Average]:::red -->|All_1| sub2[Subtract]:::red -->|All_N| pow@{ label: "Power (X<sup>2</sup>)" }
pow -->|All_1| out2[Save Channels]
in3[Channels]
in3 -->|All_C| sub3
in3 -->|All_C| p3@{ label: "Select Frame\n(Previous)"} -->|All_P| sub3[Subtract]:::red -->|All_C| out3[Save Channels]
classDef red stroke:#d88
class p3 red
See also ND processing workflows
Segmentation
Segmentation-only graphs create a Binary layer over the original image. Therefore the
graphs typically save the original All channels and the binaries from the segmentation.
In the example below two Channels are segmented:
- Threshold on
DAPIproducesNucbinary layer with segmented nuclei. - Bright Spots on
Alexa 488producesSpotsbinary layer.
---
config:
look: handDrawn
theme: neutral
---
graph TD
subgraph Output
outchn[Save Channels]
outbin[Save Binaries]
end
inp[Channels] -->|DAPI| seg[Threshold]:::red -->|Nuc| outbin
inp[Channels] -->|Alexa 488| seg2[Bright Spots]:::red -->|Spots| outbin
inp -->|All| outchn
classDef red stroke:#d88
See also Object counting workflows.
Field measurements
It measures field features such as channel intensity or ratio of channels. The measurement produces a table with one row per field (typically frame).
| Time [s] | MeanOfDAPI | MeanOfFITC | MaxOfDAPI | MaxOfFITC | MinOfDAPI | MinOfFITC |
|---|---|---|---|---|---|---|
| 2.55908 | 304.988 | 210.585 | 3,459 | 4,095 | 0 | 0 |
In the example below there is a Measure field that has an optional mask input such as ROI that is drawn interactively when the graph is ran.
---
config:
look: handDrawn
theme: neutral
---
graph TD
subgraph Output
outchn[Save Channels]
outtab[Save Tables]
end
inp[Channels] -->|DAPI| meas[Measure field]:::red -->|Records| outtab
inp[Channels] -->|All| draw[Draw ROI] -->|Mask| meas
inp -->|All| outchn
subgraph Output
end
classDef red stroke:#d88
style draw stroke-dasharray: 5 5
See also Time measurement workflows.
Object measurements
It measures object features such as size, shape, position or channel intensity. The measurement produces a table with one row per object. The main input of this measurement node is a binary layer and an optional channel for intensity measurement under the binary.
| ObjectId | Area [µm²] | Perimeter [µm] | Circularity | Elongation | MeanIntensity |
|---|---|---|---|---|---|
| 1 | 23.276 | 21.631 | 0.625 | 2.291 | 14.232 |
| 2 | 4.820 | 12.939 | 0.362 | 6.251 | 20.102 |
| 3 | 9.165 | 12.042 | 0.794 | 1.584 | 26.680 |
| 4 | 16.886 | 16.077 | 0.821 | 1.343 | 10.551 |
In the example below the Measure objects
is connected the Obj binary coming from the Threshold
and to the All channels for intensity measurement. The original channels, thresholded binaries and the
object table are all saved.
---
config:
look: handDrawn
theme: neutral
---
graph TD
subgraph Output
outchn[Save Channels]
outbin[Save Binaries]
outtab[Save Tables]
end
inp[Channels] -.->|All| meas[Measure objects]:::red -->|Records| outtab
inp[Channels] -->|DAPI| seg[Threshold] -->|Obj| meas
seg -->|Obj| outbin
inp -->|All| outchn
classDef red stroke:#d88
See also Object counting workflows.
Detecting and measuring cells
In many cases where cells must be segmented it is practical to segment nuclei first (as it is easier) and grow them following intensity (using watershed algorithm) until background intensity is met or another cell is encountered. This technique ensures relatively correct number of cells with more or less adequate cell shape.
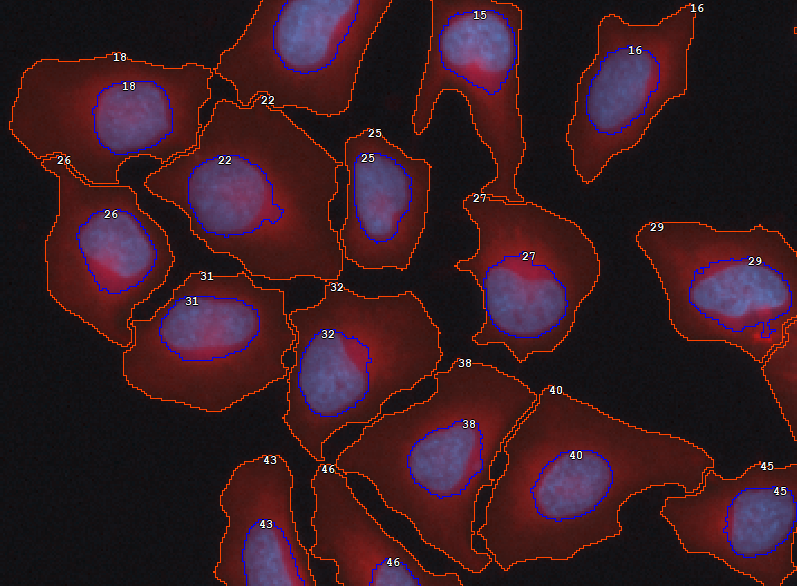
Then, to ensure that all cells are well-formed (i.e. each has one nucleus and non-empty cytoplasm) use the Make Cell node before Cell measurement.
---
config:
look: handDrawn
theme: neutral
---
graph TD
subgraph Output
outchn[Save Channels]
outbin[Save Binaries]
outtab[Save Tables]
end
inp[Channels]
seg[Threshold]:::green
grow[Grow Regions]:::green
rem[Remove on border]:::green
make[Make Cell]:::red
inp -->|DAPI| seg -->|Nuc| grow -->|Cell| rem
inp -->|DeepRed| grow
rem -->|Cell| make -->|Nuc| meas[Measure cell]:::red -->|Records| outtab
make -->|Cell| meas
inp -.->|All| meas
seg -->|Nuc| make -->|Nuc| outbin
make -->|Cell| outbin
inp -->|All| outchn
classDef red stroke:#d88
classDef green stroke:#8d8
style rem stroke-dasharray: 5 5
See also Cell measurement workflow.
Presentation of results
Records are the products of measurements. They represent tabular data. Measurements produce records one frame at a time. In some cases the task requires bigger chunk of data at once. Hence the rows have to be accumulated over a loop or overall.
In order to display the records nicely as a table or graph chart it requires making a appropriate table or chart out of the mere records.
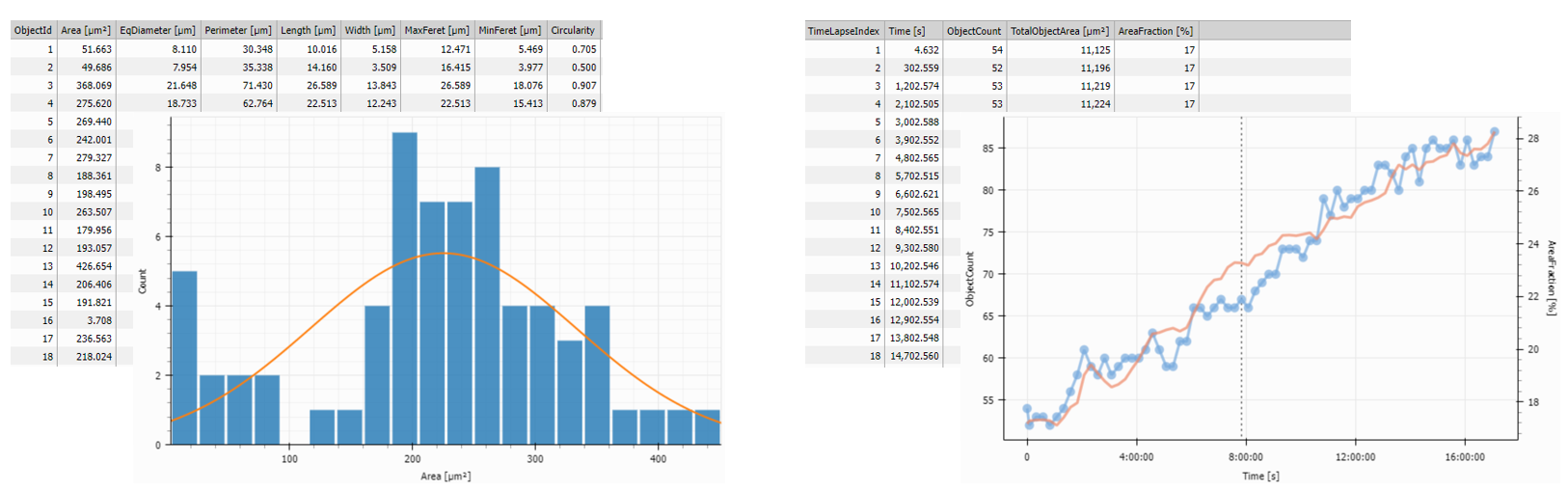
It is common to show the Tables and graphs organized near the image inside panes that allow for showing and hiding the elements as needed.
In the example below the Records output of the Object Count
measurement is accumulated using
Accumulate Records and visualized
in a Table and a Timechart showing the evolution of the number of objects over time.
The ObjTable and the Histo both contain frame data and are displayed in the left
Stacked Layout one over
the other whereas the AccumTable and the Chart containing the accumulated data are displayed in
the right Stacked Layout.
---
config:
look: handDrawn
theme: neutral
---
graph TD
subgraph Presentation
left["Stacked Layout (left)"]:::green
right["Stacked Layout (right)"]:::green
disp[Display]:::green
end
table1[Table 1]:::red
table2[Table 2]:::red
histo[Histogram]:::red
line[Line Chart]:::red
inp[Measure Objects] -->|Records| table1 -->|ObjTable| left
inp -->|Records| histo -->|Histo| left
inp2[Object Count] -->|Records| accum[Accumulate Records]:::red
accum -->|Records| table2 -->|CountTable| right
accum -->|Records| line -->|TimeChart| right
left --> disp
right --> disp
classDef red stroke:#d88
classDef green stroke:#8d8
style accum stroke-dasharray: 5 5
Tracking objects
Tracking objects connects the ‘same’ objects in subsequent frames in time.
As objects appear and disappear on each frame due to nature of the sample and segmentation imperfections the ‘same’ object typically has a different object ID on the next frame. Tracking solves this task by assigning each object a track ID which is the same for ‘same objects’.
The example below builds on object measurement. It combines track ID coming from the Track Objects with the measurement columns using Append Columns (both tables have same rows – objects). The Accumulate Tracks puts together all the frames and organizes the Records by tracks.
---
config:
look: handDrawn
theme: neutral
---
graph TD
subgraph Output
outchn[Save Channels]
outbin[Save Binaries]
outtab[Save Tables]
end
inp[Channels] -.->|All| meas[Measure objects]
inp[Channels] -->|DAPI| seg[Threshold] -->|Obj| meas -->|Records| append[Append Columns]
seg -->|Obj| outbin
inp -->|All| outchn
seg -->|Obj| tracking[Track Objects] -->|Records| append[Append Columns]
append --> accum[Accumulate Tracks] -->|Records| outtab
classDef red stroke:#d88
class tracking,append,accum red
See also Tracking workflow.
Tracking particles
Tracking particles solves the same task of assigning ID to the ‘same’ objects. But it also measures motion features like speed.
In the example below, the Time and Center measures time and position for every object, Track Particles assigns the track ID, Accumulate Tracks puts all particles into one recordset, and Motion Features calculates the dynamics of every object.
---
config:
look: handDrawn
theme: neutral
---
graph TD
subgraph Output
outchn[Save Channels]
outbin[Save Binaries]
outtab[Save Tables]
end
inp[Channels] -->|DAPI| seg[Spot Detection] -->|Spots| meas[Time and Center]
seg -->|Spot| outbin
inp -->|All| outchn
meas -->|Spot| tracking[Track Particles] -->|Records| accum[Accumulate Tracks]
accum -->|Records| mf[Motion Features] -->|Records| outtab
classDef red stroke:#d88
class meas,tracking,accum,mf red
See also Tracking workflow.