Getting started
Go to the Recipe page where the GA3 Recipes can be created and edited.
Example recipes with images
First, browse and explore Examples in the recipe page:
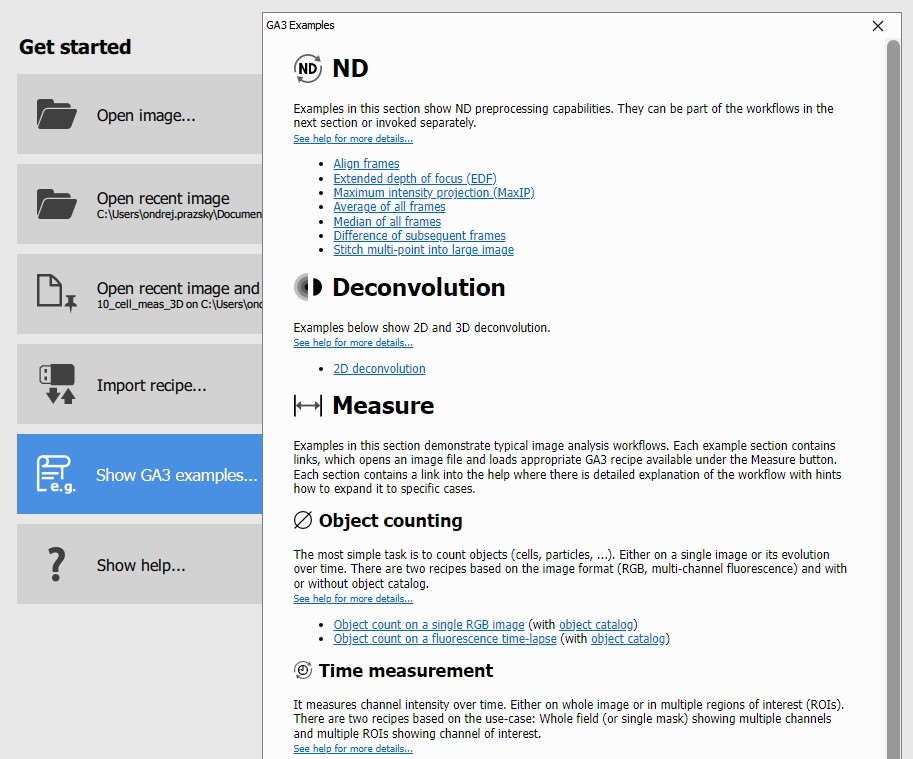
Clicking on any link will open an image with appropriate recipe to demonstrate the Workflow.
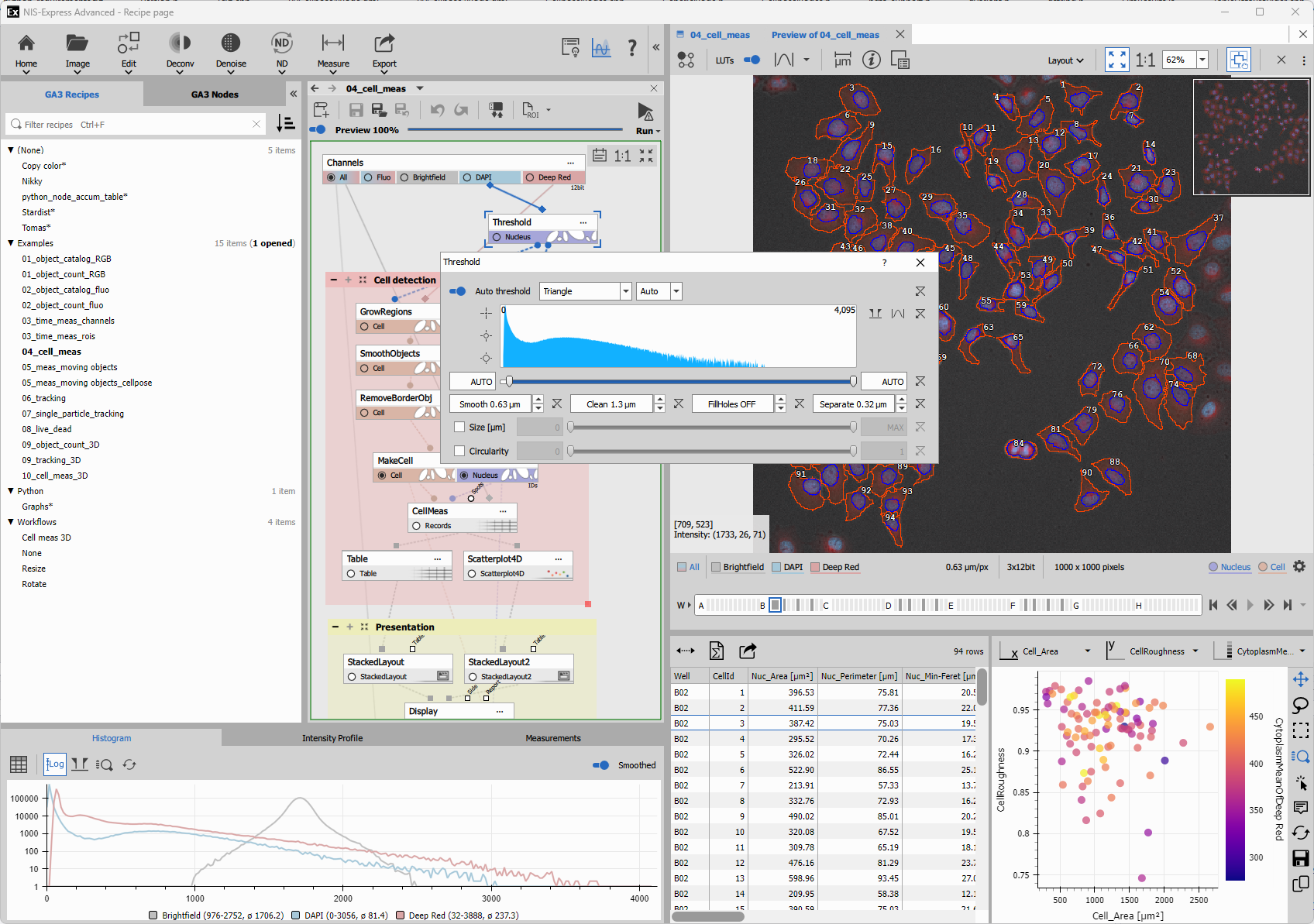
Open the workflow documentation and modify relevant nodes. Many workflow have hints on how to continue and expand the example.
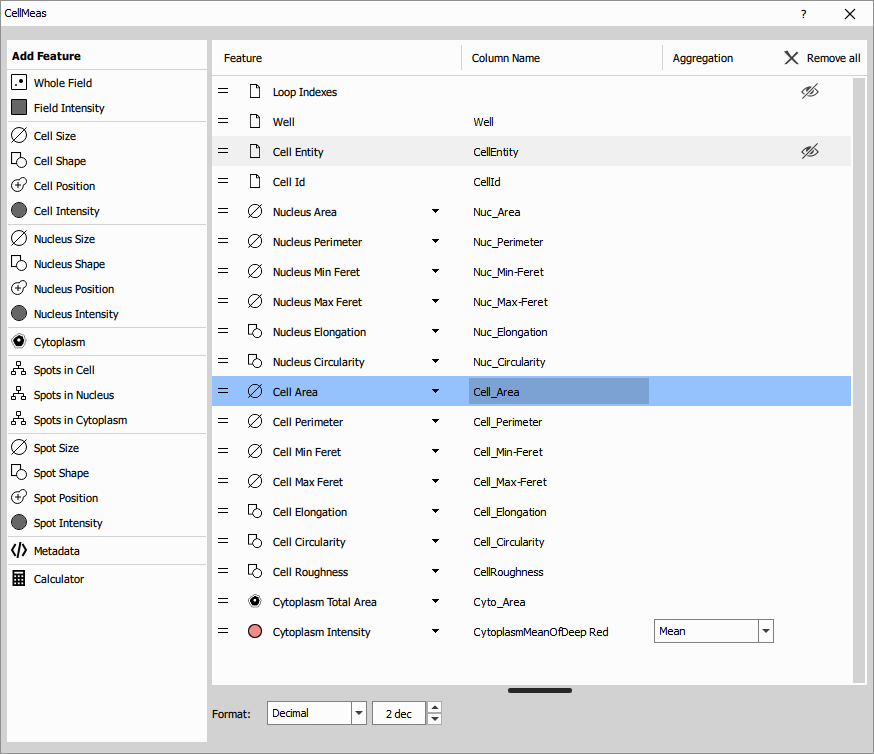
Save the modified recipe under different name. Open the original one and explore different route.
Open or Import images
Open your own images using the Image button on the Image page.
To open existing ND2 or TIFF files:
- Click Image and browse to your file
- Use Recent images or Recent folders for quick access
- Images appear in the list on the left
To import image sequences or other formats:
- Click the dropdown next to Image
- Select Import image or sequence
- Follow the import workflow to convert TIFF series, PNG, or JPEG files into ND2
Once opened, images display in the main window with their channels and overlays.
Run different recipes and compare the results
Test different analysis approaches on the same image:
- On the Image page, click the Run button
- Select a recipe from the list (e.g., “Object count” or “Cell measurement”)
- The recipe executes and results appear in the list as:
Image [Recipe name] - Click different result entries to compare segmentation and measurements
Results are saved alongside your image as .h5 files and can be reopened anytime.
Train and run custom AI
Train custom AI networks to segment your specific samples:
- Go to the AI page (Alt+A)
- Select your network type:
- Segment Objects.ai - for individual cells, nuclei, or particles
- Segment.ai - for regions or semantic segmentation
- Enhance.ai - to improve image quality
- Convert.ai - to transform between modalities
- Add your training images with annotated masks
- Set iteration count (start with 500 for standard datasets)
- Click Train and wait for completion
After training, run your network from the NIS.ai Explorer or use it in GA3 recipes. See the AI page documentation for detailed annotation guidelines.
Explore python node capabilities
Extend GA3 with custom Python code:
- Add a Python node from the ND Processing & conversions category
- Write custom processing using NumPy, scikit-image, or other scientific Python libraries
- Access the Python integration examples for:
- Custom thresholding
- Advanced segmentation with scikit-image
- Custom visualizations with Matplotlib
- Shading correction with BaSiCPy
Python nodes integrate seamlessly with standard GA3 nodes, allowing you to combine built-in and custom processing in the same recipe.